This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
trusted source
proofread
Study unveils mysterious nature of RNA editing in plants
RNA (ribonucleic acid) editing is an important process for maintaining essential functions of encoded proteins at the RNA level. Recent studies show that RNA editing is a widespread phenomenon that occurs in various land plants, including the liverworts, mosses, hornworts, lycopods, ferns, and flowering plants. However, there have been few comparative studies of chloroplast RNA editing within and between groups except for ferns.
To gain a better understanding of RNA editing in plant chloroplasts, researchers from the Wuhan Botanical Garden of the Chinese Academy of Sciences selected diverse plant species representing three distant evolutionary clades (fern, gymnosperm, and angiosperm), and determined thousands of editing sites based on the amount of RNA-seq data. The study has been published in Plant Systematics and Evolution, titled "Diversity of RNA editing in chloroplast transcripts across three main plant clades."
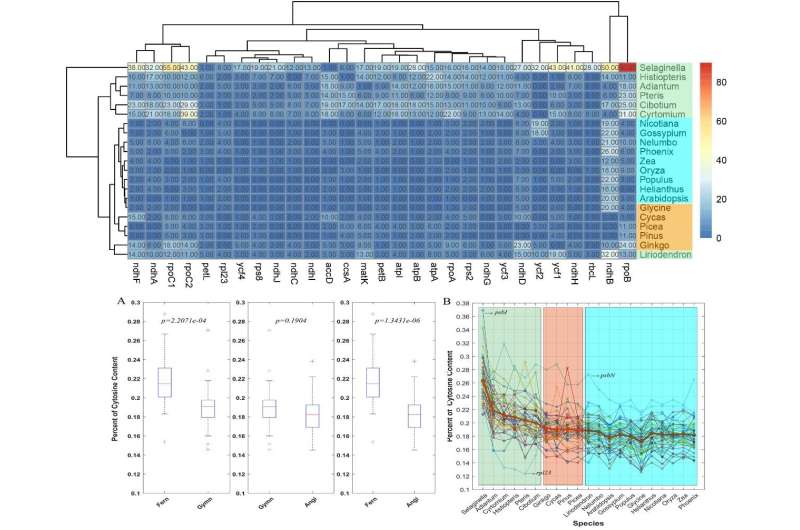
Results reveal a total of 5,203 editing sites located in chloroplast genes of 21 species after rigorous screening. The clustering relationships of the numbers of RNA editing sites roughly match the phylogenetic tree based on gene sequences, confirming that RNA editing is comparatively conservative across the plant kingdom and roughly follows the laws of evolution.
The number of editing sites decreases as plants evolve, and editing events occur more frequently in early diverging plants than in later diverging plants for each clade, suggesting that RNA editing may have occurred simultaneously in early diverging plants from different clades and suffered great loss during evolution.
In addition, the distribution of editing levels among three aspects (codon positions, species, and edited genes) was also explored. It was found that the average RNA editing level varies widely across 21 species as well as genes, with the lowest being 0.42 in Selaginella moellendorffii, and the highest up to 0.88 in the atpA gene (ATP synthase alpha-subunit gene). The decrease in cytosine content with evolution detected in this study further indicates that the situation of genomic sequence is the significant driver of editing loss in later-branching plants.
These results reveal the diversity of RNA editing in chloroplast transcripts across three major plant clades, many of the identified sites in the study have not been previously reported, and it will help to understand the enigmatic nature of RNA editing in plants.
More information: Aidi Zhang et al, Diversity of RNA editing in chloroplast transcripts across three main plant clades, Plant Systematics and Evolution (2023). DOI: 10.1007/s00606-023-01849-z
Provided by Chinese Academy of Sciences