This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Researchers develop method for deciphering positional rules in splicing
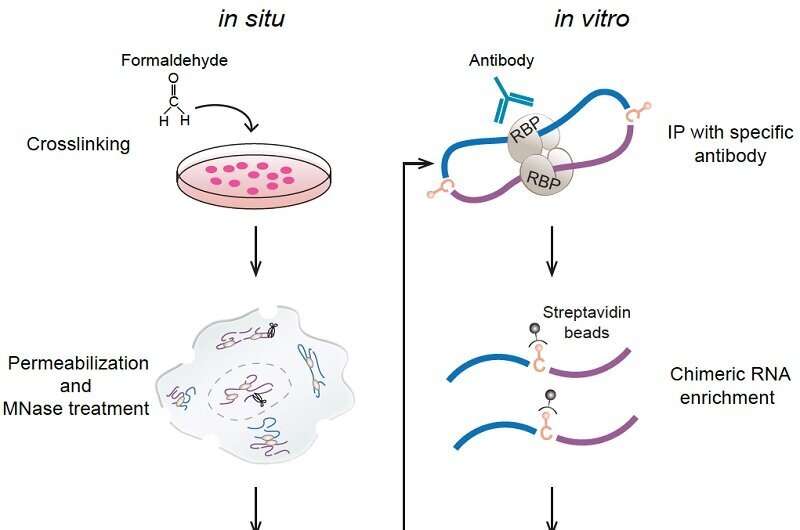
A research team led by Prof. Xue Yuanchao from the Institute of Biophysics of the Chinese Academy of Sciences has developed a new method for global profiling of in-situ RNA–RNA contacts associated with a specific RNA-binding protein (RBP) and revealed positional mechanisms by which PTBP1-associated RNA loops regulate cassette exon splicing.
This study was published online in Molecular Cell on March 22.
In eukaryotes, the same pre-mRNA can produce multiple protein isoforms to execute similar or different biological functions through alternative splicing. Several longstanding models proposed that RBPs may regulate alternative splicing by modulating long-range RNA–RNA interactions (RRI). However, direct experimental evidence was lacking.
To fill the knowledge gap, the researchers created a capture RIC-seq (CRIC-seq) method to map specific RBP-associated RRIs by leveraging the principle of RIC-seq technology and immunoprecipitation-mediated enrichment. RIC-seq technology had been developed by Prof. Xue's group in 2020 for global profiling of all expressing RBP-mediated RRIs rather than a specific one.
Using the CRIC-seq method and their self-developed data analysis pipeline, the researchers obtained high-confidence RRIs mediated by PTBP1, hnRNPA1, or SRSF1 and generated functional 3D RNA maps to investigate the positional mechanisms of these RBPs from the new angle of RNA spatial conformation.
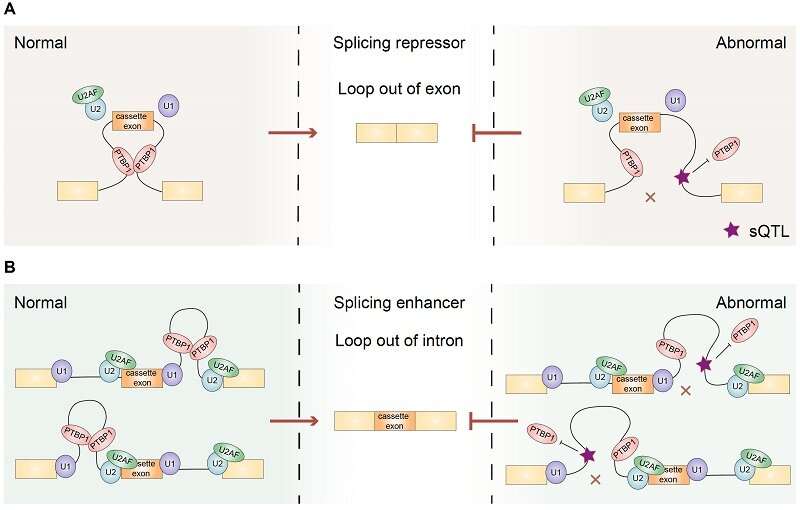
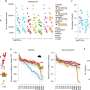
Similar to hnRNPA1, but unlike SRSF1, the researchers found that PTBP1-mediated RNA loops tended to be enriched on one side of the intron when enhancing the splicing of cassette exons. In contrast, when repressing splicing, PTBP1-mediated RNA loops tended to span the cassette exons. These findings demonstrate that PTBP1 can regulate cassette exon splicing via mediating specific RNA loops in a position-dependent manner.
According to minigene-based analysis, the researchers found that only wild-type (WT) PTBP1, but not C23S mutant PTBP1, can form a homodimer to rescue the corresponding splicing changes after PTBP1 knockdown.
Furthermore, the CRISPR-dCasRx-mediated tethering assay confirmed that RNA loops engineered by WT PTBP1 can modulate the usage of non-PTBP1-regulated cassette exons. These results demonstrate that PTBP1 dimerization can drive RNA looping to modulate splicing.
In addition, by integrating cancer-related splicing quantitative trait loci (sQTLs) with PTBP1-associated RRIs, the researchers identified 121 sQTLs that potentially affect splicing by altering RNA loops. Unexpectedly, one sQTL located in the intronic loops of HERPUD2 can induce exon 7 skipping, thus generating a short isoform to promote the proliferation of HeLa cells. These results demonstrate how cancer-related non-coding mutations modulate RNA spatial conformation to promote tumor cell growth.
More information: Rong Ye et al, Capture RIC-seq reveals positional rules of PTBP1-associated RNA loops in splicing regulation, Molecular Cell (2023). DOI: 10.1016/j.molcel.2023.03.001
Journal information: Molecular Cell
Provided by Chinese Academy of Sciences