Keeping bacteria under lock and key
Scientists and engineers are constantly looking for ways to better our world.
Synthetic biology is an emerging field with promise for improving our ability to manufacture chemicals, develop therapeutic medicines such as biopharmaceuticals and vaccines, and enhance agricultural production, among other things. It relies on taking natural or engineered pieces of DNA and combining them in new ways in biological systems, such as microbes, bacteria or other organisms.
According to University of Delaware's Aditya Kunjapur, assistant professor of chemical and biomolecular engineering, as these sophisticated microbial technologies are advanced, scientists need to explore ways to keep these organisms from ending up in the wrong environment.
For example, a bacterium that is good at making large amounts of chemicals is great in a bioreactor, but it isn't necessarily what we would want in the environment or in our food. Similarly, a specialized bacterium developed to help a plant be more productive is useful in agriculture, but we might not want that same bacterium to end up in our bodies.
Engineering traits directly into the microbe to create built-in safeguards to control where it can grow, known as biological containment, is one promising solution.
"Having the ability to control where microbes grow will help create technologies to treat disease and clean up the environment more safely," said Kunjapur, an emerging leader in biosecurity with expertise in teaching cells to create and harness chemical building blocks not found in nature.
Now, Kunjapur is the lead author of a new paper published in Science Advances on Friday, July 2 that describes progress on the stability of a biocontainment strategy, first reported in 2015, that uses a microbe's dependence on a synthetic nutrient to keep it contained. The work was done in collaboration with Harvard Medical School (HMS) and includes co-authors now at HMS, UD, Johns Hopkins University and industry.
Under lock and key
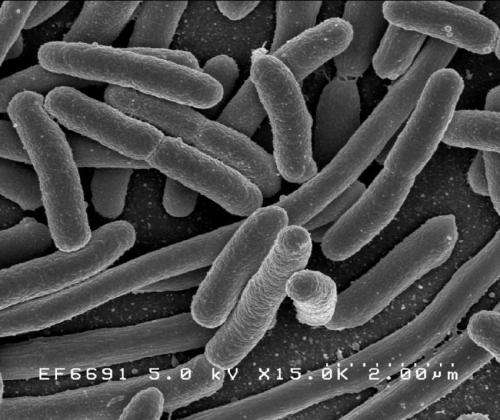
In the paper, the research team studied a strain of Escherichia coli (E. coli) that was made to depend on a synthetic nutrient for its survival, a method known as synthetic auxotrophy. They found that the strain remained stable and maintained this dependence when grown for 100 days.
"You can take enzymes that the bacteria need to grow and make them depend on a manmade building block, something it will never see in nature," said Kunjapur.
The work builds on previous research in the Church lab at Harvard Medical School, where most of this research was conducted and where Kunjapur completed postdoctoral work. In that study, the research team showed that if one enzyme required this synthetic building block, bacteria would quickly find a way out. But if three enzymes required the synthetic building block, the bacteria couldn't "unlock the cell door" within a 24-hour period to make a quick getaway.

"Asking the microbe to mutate its genome in precisely three different ways at the same time is too much, and it won't find that solution," said Kunjapur. "So, while the same 'key' might open all of the doors, the microbe can't be at all three doors at once to open them, so it remains contained."
The question was whether they would find a way out if given more time to grow.
In this new work, as the research team extended the study and grew the E. coli strain, nicknamed DEP, continuously for 100 days, they also applied extra stress by shrinking the amount of synthetic nutrient the bacteria received over time.
In three parallel experiments, the researchers found that the DEP E. coli was unable to find a way to "escape" or grow outside the cell. It did, however, adapt to a lower concentration—10 times less—of the synthetic nutrient it needed to survive. Further, the research team found that proteins on the host cell's surface changed, which had not been seen before.
"We think that as we were decreasing the amount of the nutrients, the DEP E. coli modified the proteins on the surface of the cell that regulate how much synthetic nutrient is allowed inside in order to retain more nutrients inside and keep growing," said Kunjapur. "We were surprised because at first we didn't expect them to adapt, and then we didn't expect them to adapt in this way."
To validate their results, the team went back and engineered those same genetic traits into the original ancestor E. coli strain and discovered it was able to grow at the lower synthetic nutrient concentration from the start. This information could prove useful for scaling the experiment in the future, Kunjapur said.
In a follow-up experiment, Kunjapur's colleague Eriona Hysolli, a postdoctoral fellow in the Church lab, added the bacteria directly to mammalian cells. The results were intriguing. While the bacteria grew tremendously in the first few days, over the course of a week the bacteria just faded away, without the need for antibiotics.
"Our data show the first use of recoded bacterial strains with controllable growth in a mammalian cellular model. We are excited about the promising applications in human health like the gut microbiome," said Hysolli, a co-author on the paper.
Building on this work, Michaela Jones, a UD doctoral student in Kunjapur's lab and paper co-author, conducted additional experiments that showed the bacteria stopped growing when the synthetic nutrient was taken away, then resumed growth when the nutrient was added back. Kunjapur said these findings equip researchers with new tools to control how and when bacteria grow by regulating the nutrients they need to survive.
"Now, if you need bacteria to grow at a certain rate or time, we have a synthetic building block as a lever or knob to control that," he said.
In future work, Kunjapur plans to study the biocontained DEP E. coli strain, as well as other microbe designs, under a broader set of conditions to determine if the approach will hold up in the environment. Additionally, he is working on biomedical applications for live bacterial strains, which could be used as therapeutics, probiotics and vaccines.
More information: Aditya M. Kunjapur et al, Synthetic auxotrophy remains stable after continuous evolution and in coculture with mammalian cells, Science Advances (2021). DOI: 10.1126/sciadv.abf5851
Journal information: Science Advances
Provided by University of Delaware