January 6, 2021 report
In situ sequencing of the fully structured genome

There is a sense in which the information encoded in a gene sequence can be represented by two bits per base pair location. The reality, however, is that this is far from a complete description. Although many academically and medically interesting things might be done from the minimalist sequence data, no real organism is going self-construct in developmental real time at this data rate no matter how much parallelism is used.
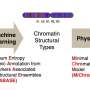
There are many reasons that this is the case. For one, the base pair symbol set is not four characters, it is literally dozens. Epigenetic modifications like methylation and acetylation change the essential character of the bases themselves, and also that of the chromatin backbone to which the bases are bound. The characters of the chromatin scaffolding itself are actively reconfigured throughout the life cycle of cell and organism. For example, during the formation of male germ cells, histones are shed and protamines temporarily swapped in. There are additionally several periods of epigenetic reboot in the germ cells of both sexes, and also in the developing embryo. These epigenetic marks are not simply on-off tags, but rather structured processes that unfold in time across dynamic genomes.
Lurking astride all this symbol logic is the big elephant in the room, namely, the spatial configuration of the DNA, and for that matter, the RNA, too. It is only now, with the advent of new ways to peer inside the nucleus, that we can begin to harvest this structural information. On the last day of the past year, researchers from the Broad Institute dropped one final bomb into the pages of Science magazine. Their paper details a new way to capture not only raw sequence information, but where in space it resides at a particular instant in time.
For now, no one is sequencing whole genomes this way. But by sequencing and localizing enough small portions of any given chromosome, a precise map of the layout of that chromosome in the nucleus can be divined. In other words, the (ACGT) bits now have (X-Y-Z) bits. This is important information to have, because different things happen in different parts of the nucleus depending if you are near the membrane, near the core, the nucleolus or other chromosomes. In no small sense, it is the configuration of a chromosome that determines which parts are accessible for transcription or replication. If one prefers, the spatial configuration can be considered as additional form of epigenetic information unto itself.
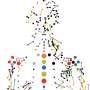
The way it all works is that instead of extracting the DNA out for sequencing, as is typically done, the researchers constructed sequencing libraries within fixed nuclei of cells. In the case at hand, either human fibroblasts or two-cell- and four-cell-stage mouse embryos. Then, they used in situ sequencing to read out the libraries via fluorescence microscopy. Each spot is essentially an amplified DNA fragment with the colors indicating the four DNA bases. A fresh round of sequencing generates each successive frame.
After the Illumina sequencing, integration across unique barcodes yields approximately 1000 spatially resolved reads per nucleus. Each of these locations includes a few hundred base-pair stretches of sequence data. This is enough to computationally connect the dots and generate nice maps of the entire nucleus to infer how the genome is folded at that moment in time. The researchers also leveraged genotype information to compare maternal and paternal chromosome organization across developmental stages.
I asked corresponding author Jason Buenrostro if their methods could be used to sequence and image mitochondrial DNA. This would be interesting for neurobiologists that want to look at things like mitochondrial heteroplasmy and selection within the spatially extended axonal and dendritic trees of neurons. He said that they haven't optimized the parameters for this yet, but it sounds like something that should be possible. One thing to be mindful of here would be how to deal with so-called NUMTs ("nu-mites") for any mitochondria that might spatially overlap with regions near the nucleus where NUMTs are found. These sequences are essentially old remnants of bygone integration of mtDNA into the nuclear genome.
One recent study found that because of unique confounding mega-NUMT variants in most of the individual trios they looked at, paternal inheritance of heteroplasmic mtDNA might be even rarer than has been previously supposed. Trios are simply the proband's genetic data (the person of interest in a sequencing study) together with the data of both parents. The output of a trio study is the phase information—i.e., the parental origin of each gene in the proband.
Since the parental genomes maintain some significant distinction in the nuclei during early stages of development, in-situ sequencing may be useful to find out how long this phenomenon persists. Quite interestingly, a new study describing fully phased human genome assembly without parental data using single-cell strand sequencing and long reads has also just been published.
Jason expects that with the foundations now laid, we should be seeing steady development of higher-resolution approaches moving forward. Anyone who wants to play around with the data in more detail can do so over at Jason's interactive lab page.
More information: In situ genome sequencing resolves DNA sequence and structure in intact biological samples. Science. DOI: 10.1126/science.aay3446 , science.sciencemag.org/content … 2/29/science.aay3446
Journal information: Science
© 2021 Science X Network