Scientists identify new methylation site on P53 guided by structure of MLL2 complex

Lysine methylation is one of the most important covalent modifications involved in diverse cellular processes. Methylation of histone H3 Lys4 (H3K4), predominantly associated with actively transcribed genes, is mainly mediated by KMT2-family histone lysine methyltransferases (KMTs). The mammalian KMT2 family contains six members (MLL1–MLL4, SET1A and SET1B), each of which has crucial and non-redundant roles in cells.
Although histone is considered the primary substrate of KMT2 family members, accumulating evidence indicates that the KMT2 family methyltransferase could also methylate non-histone substrates. However, the search for non-histone substrates of MLL family proteins has just begun.
In a study published in Structure on July 21, a research team led by Dr. Chen Yong from the Center for Excellence in Molecular Cell Science, Shanghai Institute of Biochemistry and Cell Biology of the Chinese Academy of Sciences, identified P53 as a non-histone substrate for KMT2 family methyltransferases guided by the structure of MLL2 complex.
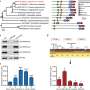
The researchers initially set out to solve the structure of MLL2-RBBP5-ASH2L complex. They found there were two M2RA complexes in one asymmetric unit induced by crystal packing. The N-terminal flanking loop (NFL) of one MLL2 extends into the substrate-binding pocket of another MLL2.
Interestingly, the configuration of MLL2 NFL is very similar to the H3 peptide as seen in the M3RA-H3 structure, but MLL2 NFL has a different sequence pattern compared with the H3 tail, indicating that KMT2 family methyltransferases could recognize and catalyze other substrates besides the canonical H3K4 sequence.
Inspired by this idea, the researchers used the ScanProsite tool to search the possible non-histone human substrates containing a similar sequence as MLL2 NFL, and then identified P53 as one of the top candidates.
Subsequently, through a series of biochemical experiments and mass spectrometry analysis, they demonstrated that K305 of P53 could be methylated by KMT2 family complexes, expanding the landscape of non-histone substrates for KMT2 family methyltransferases.
P53 contains four known lysine methylation sites which are all located on the C-terminal tail. The newly-identified methylation site in this study is positioned in a loop region between the DNA-binding and tetramerization domains, which may affect the transcription activity of P53.
This study not only provides insights into the crosstalk between KMT2-family proteins and P53, but also inspires further studies of KMT2-mediated non-histone methylation in normal physiological functions and oncogenic processes.
More information: Yanjing Li et al. Crystal Structure of MLL2 Complex Guides the Identification of a Methylation Site on P53 Catalyzed by KMT2 Family Methyltransferases, Structure (2020). DOI: 10.1016/j.str.2020.07.002
Journal information: Structure
Provided by Chinese Academy of Sciences