New X-ray imaging approach could boost nanoscale resolution for advanced photon source upgrade
A longstanding problem in optics holds that an improved resolution in imaging is offset by a loss in the depth of focus. Now, scientists are joining computation with X-ray imaging as they develop a new and exciting technique to bypass this limitation.
The upcoming Advanced Photon Source Upgrade (APS-U) project at Argonne will put this problem under one of the brightest spotlights imaginable. The upgrade will make the APS, a Department of Energy Office of Science User Facility, 500 times brighter than it is today, further enhancing the capabilities of its X-rays to study the arrangements of atoms and molecules in a wide range of biological and technological materials.
"A whole variety of X-ray imaging experiments ultimately will need something like this as they all push the resolution to finer length scales in the future," said Chris Jacobsen, an Argonne Distinguished Fellow and professor of physics at Northwestern University. With the Upgrade in place, the APS's X-rays could allow scientists to study systems like the brain's full network of synaptic connections, or the entire volume of an integrated circuit down to its finest details.
"A whole variety of X-ray imaging experiments ultimately will need something like this as they all push the resolution to finer length scales in the future."—Chris Jacobsen, Argonne Distinguished Fellow/professor of physics at Northwestern University.
In a new study, a team of researchers from Argonne, Northwestern and Cornell University has developed a computational approach that simulates how the APS-U might best operate, particularly in X-ray imaging mode, known as ptychography. They presented their approach, called Multislice Optimized Object Recovery (MOOR), in the Sept. 20 issue of Optica.
X-ray ptychography is a powerful scanning technique that images thin slices of materials at a resolution of better than 30 nanometers (a human hair measures approximately 75,000 nanometers in diameter). Traditional algorithmic and computational approaches used to reconstruct images from samples probed with this technique have been limited to two-dimensional slices.
"The scaling up of X-ray ptychography for 3-D brain imaging would be enormous," said Genia Kozorovitskiy, an assistant professor of neurobiology at Northwestern. "Our neural circuits are wired together by trillions of tiny junctures called synapses, which are used for cell-to-cell communications through electrochemical means. Changes in how neural circuits and synapses are wired together hold the key to understanding the cause of many human neurological and mental health disorders."
The National Institutes of Health are funding Jacobsen and Kozorovitskiy to optimize both the resolution and sample preparation of neural tissue for X-ray imaging. Increasing depth penetration of X-ray ptychography through the APS-U would enable researchers to create a new, faster, non-destructive way to map the connective neural tissue of entire vertebrate brains.
Neuroscientists today use electron microscopy to map neural connectivity in mice. "Sectioning the full mouse brain for electron microscopy is a daunting task, one that no one has yet attempted," Kozorovitskiy noted. "With X-ray tomography, the entire brain could in theory be imaged with no physical sectioning, simplifying large-scale, high-resolution image construction."
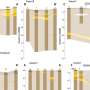
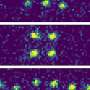
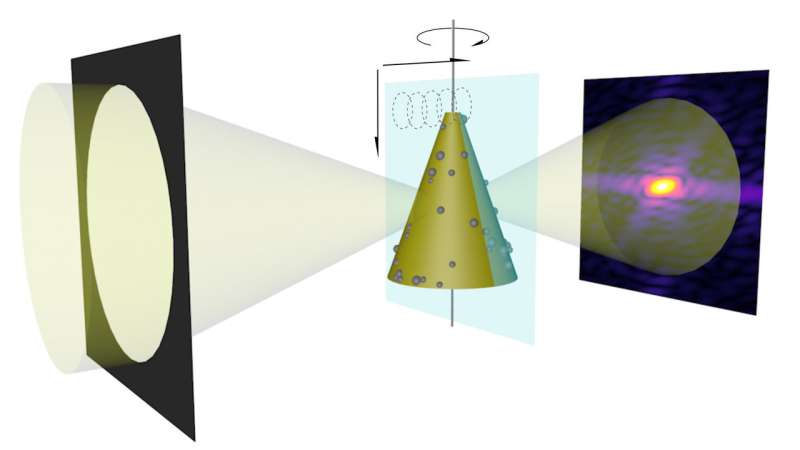
To test the effectiveness of the MOOR algorithm for this purpose, the scientists devised a test sample whose properties would elicit the depth-of-focus limit that they sought to overcome. They designed the object, a tapered glass hollow cone measuring 200 nanometers along its axis and embedded with titanium nanospheres, to resemble the thin capillary tubes that are widely used in laboratory experiments for the microscopic study of cells frozen in liquid suspension.
This simple, easily manufactured sample ensured that the first experimental measurements the scientists will take can match their model. "Our longer-term goal is not to study glass capillaries with titanium dioxide spheres on them, but to really push the resolution for imaging a whole mouse brain," Jacobsen said.
Researchers currently use the Bionanoprobe in sector 9 of the APS for 3-D imaging of small biological samples, such as a frozen, hydrated cell. Efforts are underway to develop the same capability on much larger samples, including whole mouse brains, at other instruments at the APS.
The MOOR demonstration involved solving an optimization involving nearly 17 million complex variables, with the MOOR algorithm being scaled to 2,880 cores of the Bebop supercomputing cluster at Argonne's Laboratory Computing Resource Center. The work will further benefit from a new computing award as part of the Data Science Program at the Argonne Leadership Computing Facility, another DOE Office of Science User Facility.
The relatively easy part of the problem is to understand what happens when moving from model to measurement.
"We can solve this problem by baking the physics into the model," said Youssef Nashed, a computer scientist in Argonne's Mathematics and Computer Science division. "The physics tells us how to go from model to measurement. The math tells us how to go from measurement to model. It's an inverse problem."
To solve the inverse problem, the Argonne optimization algorithm iteratively refines its representation of the unknown model in seeking agreement between the model and the massive volumes of imaging data.
"To do this efficiently at the resolution needed requires a strong mix of applied mathematics, parallel computing and physics," Nashed said.
This mixing of disciplines also pertains to applications of technique. "It's exciting that algorithms such as MOOR can serve as a numerical lens for imaging the increasingly nanoscale-level circuits that are the workhorses of modern computing hardware," said Stefan Wild, a co-author of the Optica article and a computational mathematician at Argonne.
More information: M. A. Gilles et al. 3D x-ray imaging of continuous objects beyond the depth of focus limit, Optica (2018). DOI: 10.1364/OPTICA.5.001078
Journal information: Optica
Provided by Argonne National Laboratory