The faulty yardstick in genomics studies and how to cope with it
Geneticists use standards to reconstruct the history of a species or to evaluate the impact of mutations, in the form of genetic markers scattered throughout the genome. Provided these markers are neutral, i.e. that they have evolved randomly rather than through a selective process, they can be reliably used as standards to compare various parameters across populations.
However, scientist Fanny Pouyet and colleagues from the Group of Laurent Excoffier at the SIB Swiss Institute of Bioinformatics and University of Bern, have recently discovered that 95 percent of the genome actually seems to be affected by selection and other genetic biases and that markers previously thought to be neutral appear to provide skewed estimates. Their study, published in eLife, calls for the re-examination of a plethora of results and provides the tools and recommendations to correct such issues in the future.
Models used to reconstruct the history of a species or to discover how populations are related to one another rely on a key assumption: that the genome regions under scrutiny are made of "neutral" snippets of DNA, i.e. parts that have evolved randomly rather than being selected for or against. But these regions might actually not be as neutral as previously thought, according to a recent finding by scientists at SIB and the University of Bern: "What we find is that less than 5 percent of the human genome can actually be considered as 'neutral'", says Fanny Pouyet, lead author of the study. "This is a striking finding: It means that 95 percent of the genome is indirectly influenced by functional sites, which themselves represent only 10 percent to 15 percent of the genome," she concludes. These functional sites encompass both genes and regions involved in gene regulation.
A "universal" recipe for neutral markers
Scientists have long devised the best way to obtain "unbiased" sets of genomic markers and several such sets are routinely used in genetic studies. The study of Pouyet and colleagues now sheds concern on the reliability of these markers. "We re-examined all existing sets of markers presented as "neutral" and found that they provided, under one aspect or another, skewed estimates," indicates Pouyet. The team then went on to identify a new set of markers that matched, this time, all the neutrality criteria, using two whole genome datasets of over a hundred individuals in total. This neutral dataset has now been made available for humans, but the method could in theory be used to find such markers in any other species.
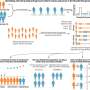
How has the use of non-neutral markers affected demographic inferences so far? In order to obtain an initial assessment of the situation, the team compared the outcomes of the use of non-neutral vs. neutral markers in the context of contemporary African and Japanese populations. "We found that such bias could lead one to wrongly infer that populations of constant size have grown, or to overlook events that drastically reduce the size of a population," Excoffier points out. "While the nature and extent of the bias is difficult to predict for a given population, one thing that is certain is that the demography of all human populations should be re-examined on the basis of the new set of neutral markers. Actually not only demography: a biased neutral reference could also affect the measure of the impact of mutations," he concludes.
More information: Fanny Pouyet et al, Background selection and biased gene conversion affect more than 95% of the human genome and bias demographic inferences, eLife (2018). DOI: 10.7554/eLife.36317
Journal information: eLife
Provided by Swiss Institute of Bioinformatics