Crowdsourcing friendly bacteria helps superbug cause infection
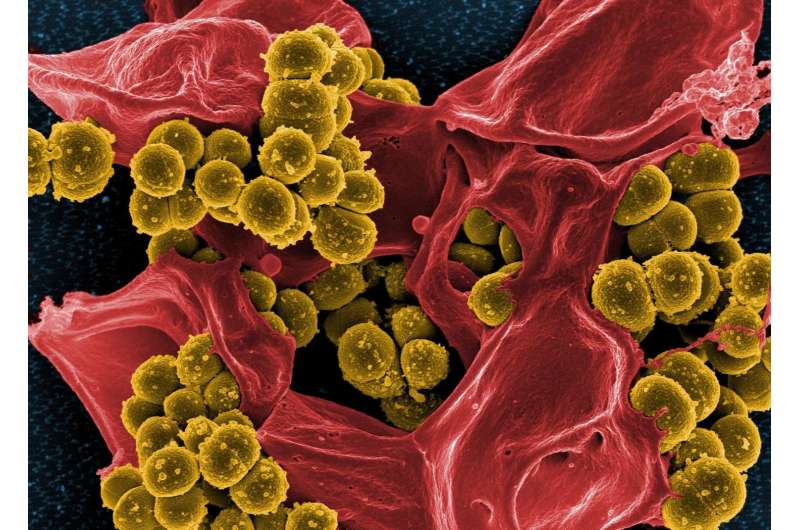
Antimicrobial resistant pathogens crowdsource friendly bacteria to survive in immune cells and cause disease, a new study by the University of Sheffield has revealed.
Scientists have discovered the human pathogen Staphylococcus aureus (S.aureus), uses benign bacteria present in the skin to initiate infection.
Known commonly as its infamous antimicrobial resistant form MRSA (meticillin-resistant Staphylococcus aureus), the ground-breaking research discovered that by using the other bacteria present on the skin, the pathogen can survive the mechanisms our immune system deploys to destroy it.
The findings, published today (16 July 2018) in Nature Microbiology, give scientists a new insight into the mechanisms of the so-called superbug, which is hard to treat due to its resistance to several widely used antibiotics.
Simon Foster, Professor of Molecular Microbiology at the University of Sheffield and lead author of the study, said: "The remarkable discovery that S.aureus uses the presence of the otherwise benign members of the human skin microflora to augment disease, changes our understanding of how pathogens can elude our defences and initiate infection.
"We harbour a vast array of bacteria as microflora in our guts and on our skin so understanding this combination of pathogen, native microflora and host provides new avenues for approaches to prevent and treat infection.
"A lot of people have been trying to develop a vaccine against the potentially life-threatening virus for a number of years without success.
"This study has shown how a potentially fatal disease like MRSA gains access to our bodies and how the organisms are able to survive inside immune cells—phagocytes."
The pioneering five-year project was led by the Florey Institute at the University of Sheffield in collaboration with an international team of scientists from Canada, Holland, Sweden and the UK.
Professor Foster added: "All infectious disease starts from an environment of mixed bacteria and other material so this might have much wider ramifications.
"The study has important implications for disease because with our work we can drop the infectious dose by over 1,000 fold.
"Importantly, S. aureus is not a dominant member of our bacterial flora and so this study can explain how relatively few pathogenic bacteria can crowdsource from the other organisms present on our skin to allow it to set an infection that may ultimately prove fatal.
"It alters the way that we view how infection occurs, the way it should be studied and sets the scene for how it might be tackled."
More information: Emma Boldock et al, Human skin commensals augment Staphylococcus aureus pathogenesis, Nature Microbiology (2018). DOI: 10.1038/s41564-018-0198-3
Journal information: Nature Microbiology
Provided by University of Sheffield