Platforms for investigating lncRNA functions
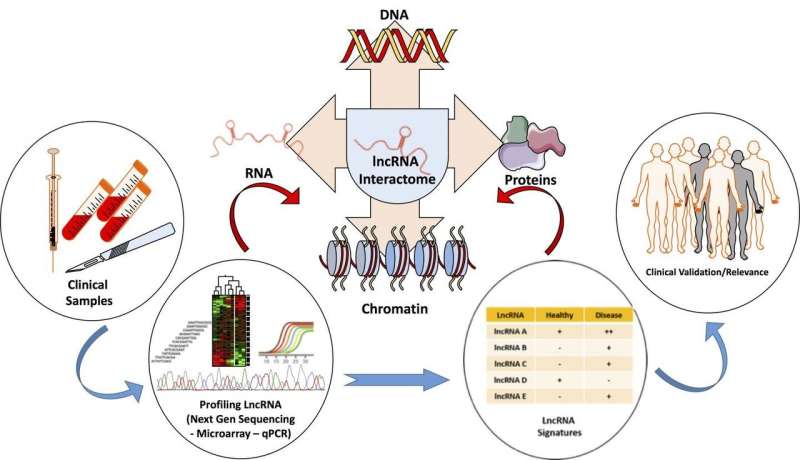
To aid in the discovery and understanding of lncRNA biology, newly published work from Richard and Eichhorn in SLAS Technology features the technological platforms and methodology presently used to identify the roles of lncRNA in biology. This work highlights the databases and tools used to study lncRNA and techniques used to study their function.
Prior to sequencing of the human genome, it was presumed that most of the DNA coded for proteins with the rest of the human genome classified as "junk DNA." However, recently these vast stretches of junk DNA have been annotated and discovered to contain thousands of non-coding RNA (ncRNA), including a family of transcripts referred to as long noncoding RNAs (lncRNAs).
Although still in its infancy, research into the biology of lncRNAs is demonstrating the importance of lncRNAs in development and disease. LncRNAs are implicated in many biological processes ranging from housekeeping functions such as transcription to more specialized functions such as dosage compensation or genomic imprinting. Furthermore, lncRNAs can regulate varied activities such as messenger RNA degradation, translation and effect protein kinetics or function by acting as RNA decoys or scaffolds. With its vast assortment of biological activity, identifying the mechanistic action of lncRNA may seem overwhelming.
More information: John Lalith Charles Richard et al, Platforms for Investigating LncRNA Functions, SLAS TECHNOLOGY: Translating Life Sciences Innovation (2018). DOI: 10.1177/2472630318780639
Provided by SLAS (Society for Laboratory Automation and Screening)