Bacteria in estuaries have genes for antibiotic resistance
An international group of researchers, including Professor Michael Gillings from Macquarie University, have reported that pollution with antibiotics and resistance genes is causing potentially dangerous changes to local bacteria in estuaries.
Polluting antibiotic agents in these waterways, they say, help the bacteria to acquire genes that make them less responsive to antibiotics – known as antibiotic resistance genes. These genes could then enter our food chain when we eat aquatic animals from these areas.
"By eating fish or crustaceans, for example, sourced from estuary regions, we could be exposed to rare antibiotic resistance genes that could spread quickly through large numbers of people. These resistance genes could change the natural bacteria in our own bodies, affecting our health in a multitude of ways," explained Professor Gillings, who studies how antibiotic pollution of waterways creates superbugs.
The researchers found that within these estuary regions around one antibiotic resistance gene existed per bacterial cell – meaning that the great majority of bacteria had at least one gene that helped them become resistant to antibiotics.
"We detected over 200 different resistance genes in total, meaning that we're dealing with quite a variety of antibiotic resistance genes that could help estuary bacteria along the path to becoming superbugs," explained Professor Gillings.
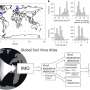
The study, which tested 4,000 km of China's coastline, found that estuaries generally had levels of antibiotic resistance gene pollution at around 1 million resistance genes per gram of estuarine sediment, and in some cases up to 100 million per gram.
"This tells us that antibiotic resistance genes are now common in bacteria in estuaries –China's estuaries are just a microcosm of a worldwide problem," said Professor Gillings.
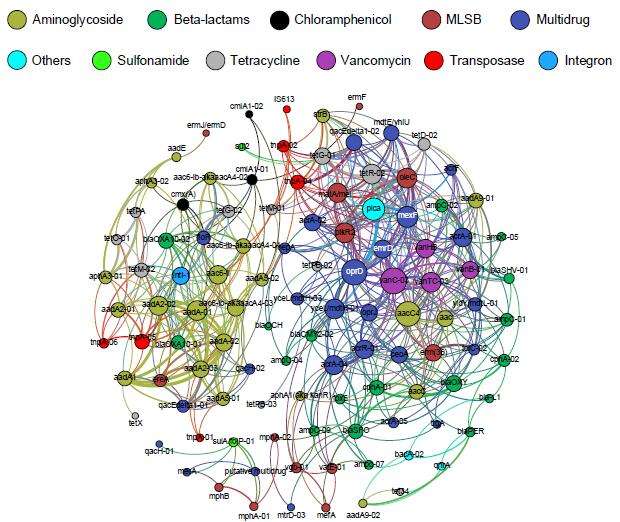
"There are plenty of famous estuarine regions in Australia, in the city of Sydney we have Port Stephens, Sydney Harbour, Botany Bay and the Hawkesbury River, which all have significant sources of microbial contamination and pollution, particularly following rainfall. These regions are places where bacteria and polluting antibiotics can meet, promoting the accumulation of antibiotic resistance genes," he concluded.
More information: Yong-Guan Zhu et al. Continental-scale pollution of estuaries with antibiotic resistance genes, Nature Microbiology (2017). DOI: 10.1038/nmicrobiol.2016.270
Journal information: Nature Microbiology
Provided by Macquarie University