Researchers sequence genome of 'gluttonous' tobacco hornworm
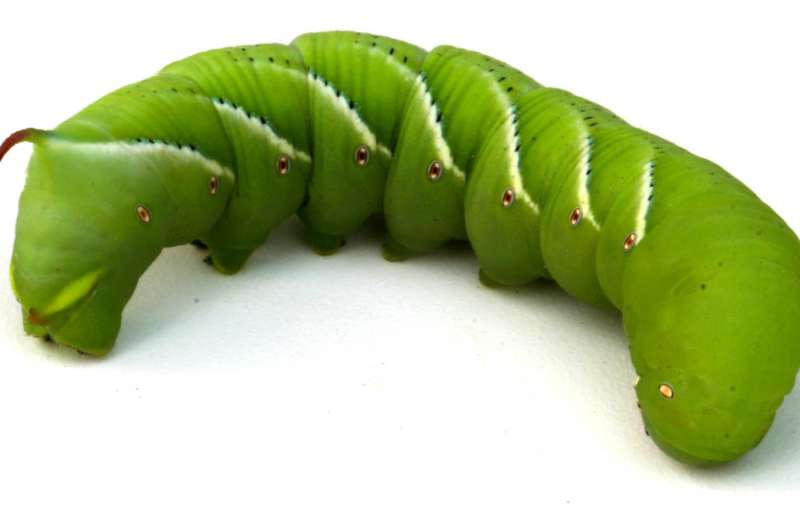

An international team of researchers has sequenced the genome of the tobacco hornworm—a caterpillar species used in many research laboratories for studies of insect biology.
Professor Gary Blissard of the Boyce Thompson Institute at Cornell University, and Professor Michael Kanost of Kansas State University, initiated the study and are co-senior authors on this large international project that included 114 researchers from 50 institutions and 11 countries.
The researchers have published their work in a paper titled "Multifaceted biological insights from a draft genome sequence of the tobacco hornworm moth, Manduca sexta" in the journal Insect Biochemistry and Molecular Biology. The scientists have made the genome sequence available to the public through the National Agricultural Library and the National Center for Biotechnology Information (NCBI).
"The completion of this project marks a major milestone in the study of insect biochemistry and molecular biology, as Manduca sexta is an important model insect: one that has been studied for its physiology and biochemistry for many decades," said Blissard.
"This project represents years of collaborative research across the world," said Kanost. "We wanted to provide these valuable data to scientists, and our hope is that this sequenced genome will stimulate new research in molecular studies of insects."
The tobacco hornworm, or Manduca sexta, develops into the Carolina sphinx moth. The name Manduca comes from the Latin word for glutton because these caterpillars eat so much. M. sexta occurs naturally in North, Central and South America and is a known pest to gardeners: It eats the leaves of tomato plants and also can be found on pepper, eggplant and potato plants. Crops and weeds from this plant family, which includes tobacco, produce chemicals such as nicotine that deter feeding by most insects, but not M. sexta, which makes its physiology especially interesting to scientists.
The sequenced genome can lead to improved molecular biology, physiology and neurobiology research in insects and also may help in developing future new methods for insect pest management. The tobacco hornworm is a good model species because of its large size—the caterpillar can measure up to 4 inches long— making it easy to collect tissue samples.
Kanost has studied the tobacco hornworm for decades, and he and Blissard decided to start the collaborative project to sequence the tobacco hornworm's genome in 2009. Collaboratively, the research team sequenced the DNA that encodes the genes as well as the RNA from the insect at different developmental stages, to identify when different genes are expressed and in which tissues and organs.
Kanost and the Kansas State University team prepared and purified the DNA of the tobacco hornworm and sent the samples to the Baylor College of Medicine Human Genome Sequencing Center in Houston, which performed the genome sequencing. Blissard's group at the Boyce Thompson Institute isolated RNA from a broad range of tissues and specific developmental stages throughout the egg, larval, pupal and adult stages of the insect. The international team used a common computer system so that researchers from around the world could analyze the gene sequences based on their areas of expertise.
Blissard studies the interactions between viruses and their insect hosts. He and his colleagues examined the vacuolar protein sorting (VPS) genes. These genes code for proteins involved in transporting small containers called vesicles throughout the healthy cell, but these proteins are often hijacked by viruses in order to move the virus through the cell.
"What makes the project unique is that so many different groups did in depth studies of so many gene families," said Blissard "and this is an exceptionally rich resource that will be useful not only to scientists studying insects, but also to scientists studying other organisms and their pathogens and diseases."
More information: Michael R. Kanost et al, Multifaceted biological insights from a draft genome sequence of the tobacco hornworm moth, Manduca sexta, Insect Biochemistry and Molecular Biology (2016). DOI: 10.1016/j.ibmb.2016.07.005
Provided by Boyce Thompson Institute










.jpg)