Visualizing and predicting evolution by mapping the elusive 'fitness landscape'
Suppose you were trying to design a vaccine to combat next season's influenza virus. Having a detailed map that tells you exactly how various strains of the flu bug will evolve would be extremely helpful.
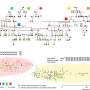
Creating just that sort of map is the goal of evolutionary biologists who study a conceptual tool called the fitness landscape, which provides a way to visualize and predict evolution.
The idea of the fitness landscape has been around since the 1930s, when it was proposed by population geneticist Sewall Wright. But mapping a detailed fitness landscape is a daunting challenge, and the landscapes created to date are fairly crude.
Now, a team of University of Michigan researchers is reporting the first comprehensive in vivo fitness landscape of a gene—roughly 100 times larger than any previous effort. The findings are expected to be of interest to evolutionary biologists, geneticists and molecular biologists, said team leader Jianzhi "George" Zhang, a professor in the U-M Department of Ecology and Evolutionary Biology.
A paper summarizing the team's findings, which are based on the manipulation of a single gene in baker's yeast, is scheduled for online publication in the journal Science on April 14.
"The concept of the fitness landscape is critically important and underlies many evolutionary theories. But until now, we were simply not able to measure it," Zhang said. "We still have a long way to go, but this is a big step toward measuring fitness landscapes."
Think of a fitness landscape as a 3-D map that allows researchers to visualize the relationship between an organism's genetic makeup and its ability to produce offspring, which biologists call fitness. Reproductive rate is the ultimate measure of evolutionary success—really the only metric that matters.
Darwinian fitness is determined by the interplay between an organism's genes and its environment. In any given environment, an organism's genetic makeup, or genotype, can either help or hurt it in the competition against other individuals of the same species.
The fitness landscape is a way of visualizing the relationship between genotypes and reproductive success. But moving the study of fitness landscapes from a theoretical pursuit to an experimental science has been challenging for several reasons. First off, the genotype space is immense, consisting of vast numbers of genetic possibilities for any single organism.
The human genome, for example, is a sequence of about 3 billion letters built from the four-letter genetic alphabet of A, T, G and C. At each position in the genetic sequence there are four letters to choose from, which means there are billions and billions and billions of possibilities in a complete human genotype.
To make this problem more manageable, Zhang and his colleagues focused on one small gene from baker's yeast, a single-celled organism with about 6,000 genes. Yeast is commonly used by scientists to provide insights about genetics and cell biology.
The researchers selected a gene that holds the genetic instructions for making transfer RNA, or tRNA, which is part of the cellular machinery used to assemble proteins from amino acids. The gene they selected contains just 72 letters known as nucleotide bases.
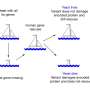
To map a fitness landscape for this tRNA gene, the researchers needed to assess the reproductive success associated with every possible combination of A, T, C and G at each of the 72 locations in that gene sequence.
So they constructed more than 65,000 yeast strains, each carrying a unique variant of the gene with one or more misspellings of the sequence. Each misspelling is called a point mutation.
The 65,000-plus yeast strains were then placed together in a test tube and allowed to grow for one day through asexual cell division. At the end of the 24-hour competition period, the growth rate of each strain—essentially a measure of its Darwinian fitness—was determined.
The result was a fitness landscape for a tRNA gene, the first relatively comprehensive, in vivo fitness landscape of any gene in any eukaryotic organism, according to Zhang.
The researchers showed that about 1 percent of the point mutations were beneficial to the organism, while 42 percent were harmful. And when two harmful mutations interacted, the outcome was usually more harmful to the organism than if the two mutations had acted independently.
The U-M team also found that fitness is broadly correlated with the predicted fraction of correctly folded tRNA molecules, revealing a biophysical basis for the fitness landscape.
"The experimental method we developed for measuring fitness landscapes can also be applied to other genes, including protein-coding genes," Zhang said. "So we expect that our paper will be of broad and immediate interest."
More information: "The fitness landscape of a tRNA gene" Science DOI: 10.1126/science.aae0568
Journal information: Science
Provided by University of Michigan