October 7, 2015 report
Genetic diversity of sub-species of a malaria-causing parasite found
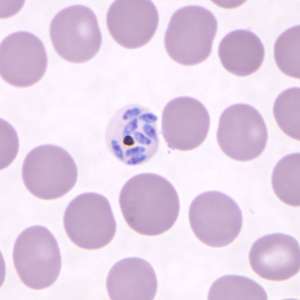
(Phys.org)—An international team of researchers has found that three sub-species of Plasmodium knowlesi, parasites that cause malaria, are genetically diverse and diverge between sub-populations. In their paper published in Proceedings of the National Academy of Sciences, the team describes their study in great detail and their opinion on whether the sub-species present a threat to human health.
Malaria is, of course, a disease that is caused by infestation of a parasite. Presently, there are two main species that cause problems for humans; Plasmodium vivax and Plasmodium falciparum—a third Plasmodium knowlesi, can cause infections in people, but it is much more often seen in long and pig-tailed macaques. Prior research has shown that two of the sub-species mainly infect macaques and that a third, of which less is known, exists in some parts of Southeast Asia. In this new effort, the research team has found that divergence among the sub-species varies greatly.
Study of P. knowlesi has begun in earnest after health officials began to worry that it was becoming more likely to infect humans as populations expanded into macaque territory—reports of travelers in some areas becoming infected with the parasite species only served to increase concerns. P. knowlesi is spread the same way as other types of the parasite, i.e. via mosquitoes. What has not been clear is whether it is able to spread between just humans through the mosquito vectors. To learn more, the researchers conducted illumina paired-end genome sequencing from nearly fifty samples collected from macaques in Borneo, comparing them to one another and to other species of the parasite, which mainly originated in the Malaysian Peninsula and the Philippines. This work resulted in the creation of profiles of the three subpopulations of P. knowlesi, which showed strong signals of natural diversity. They showed also what the researchers describe as "a remarkable substructure consisting of two major sympatric clusters of the clinical isolates and a third cluster comprising the laboratory isolates."
The good news is that the team also found that the sub-species showed little evolutionary change over time that indicated they were on a path towards becoming more transmissible to humans—most infections, they note, still occur in long-tailed macaques.
More information: Population genomic structure and adaptation in the zoonotic malaria parasite Plasmodium knowlesi, Samuel Assefa, PNAS, DOI: 10.1073/pnas.1509534112
Abstract
Malaria cases caused by the zoonotic parasite Plasmodium knowlesi are being increasingly reported throughout Southeast Asia and in travelers returning from the region. To test for evidence of signatures of selection or unusual population structure in this parasite, we surveyed genome sequence diversity in 48 clinical isolates recently sampled from Malaysian Borneo and in five lines maintained in laboratory rhesus macaques after isolation in the 1960s from Peninsular Malaysia and the Philippines. Overall genomewide nucleotide diversity (π = 6.03 × 10−3) was much higher than has been seen in worldwide samples of either of the major endemic malaria parasite species Plasmodium falciparum and Plasmodium vivax. A remarkable substructure is revealed within P. knowlesi, consisting of two major sympatric clusters of the clinical isolates and a third cluster comprising the laboratory isolates. There was deep differentiation between the two clusters of clinical isolates [mean genomewide fixation index (FST) = 0.21, with 9,293 SNPs having fixed differences of FST = 1.0]. This differentiation showed marked heterogeneity across the genome, with mean FST values of different chromosomes ranging from 0.08 to 0.34 and with further significant variation across regions within several chromosomes. Analysis of the largest cluster (cluster 1, 38 isolates) indicated long-term population growth, with negatively skewed allele frequency distributions (genomewide average Tajima's D = −1.35). Against this background there was evidence of balancing selection on particular genes, including the circumsporozoite protein (csp) gene, which had the top Tajima's D value (1.57), and scans of haplotype homozygosity implicate several genomic regions as being under recent positive selection.
Journal information: Proceedings of the National Academy of Sciences
© 2015 Phys.org