September 8, 2015 report
Understanding reef systems at the genetic level

(Phys.org)—Coral reefs are the most diverse marine ecosystems, biodiversity hotspots now under anthropogenic threat from climate change, ocean acidification and pollution. Efforts are underway to protect and expand shrinking reef systems, but such endeavors are inhibited by the lack of information about such fundamental features as the functional symbiosis between the cnidarian coral animal and the photosynthetic alga that live in its gastrodermal cells.
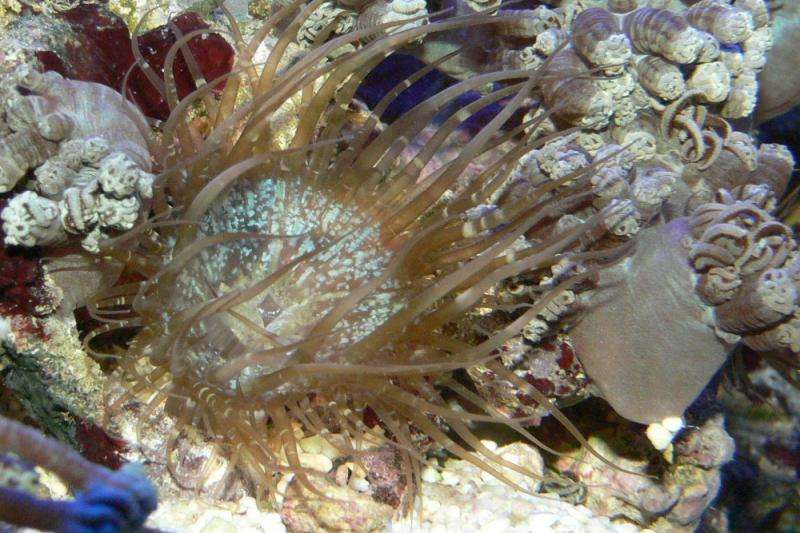
Researchers have been seeking an experimentally versatile model of the cellular biology underlying this symbiosis, which could be a key to understanding the adaptability and resilience of reef systems. An international group of researchers now reports in the Proceedings of the National Academy of Sciences on a genetic study of the sea anemone Aiptasia, a globally distributed species that harbors Symbiodinium, the most widespread known group of endosymbiotic dinoflagellates, which inhabit many species and are common to cnidarians that occupy nutrient-poor waters.
The host provides the algae with a sheltered environment and the nutrients necessary for photosynthesis and growth; in return, Symbiodinium supplies over 90 percent of the cnidarian's total energy. The reasons this holobiont provides an attractive model system for reef studies include polyp sizes convenient for experimentation, and its easy production in a laboratory environment, where it can reproduce asexually, yielding large clonal populations. It can also be maintained indefinitely in an aposymbiotic state (without its endosymbiote) and reinfected with a variety of Symbiodium strains.
The researchers sequenced the DNA from Aiptasia anemones and produced a reference transcriptome, which was sequenced using RNA derived from different developmental and symbiotic states. They uncovered a number of previously unknown genetic features of Aiptasia, but more importantly, the study provides a foundation for understanding the evolution and function of the symbiosis between the two organisms.
Among their discoveries, the researchers found a novel cnidarian-specific family of putative pattern-recognition receptors that may be involved in the symbiotic relationship. Cnidarian hosts must necessarily distinguish among potential symbionts, and most can establish relationships with some strains of Symbiodinium but not with others. "Such discriminations must be accomplished in the absence of an adaptive immune system and presumably depend on innate immunity mechanisms that involve the recognition of microbial cell-surface molecules by host pattern-recognition receptors," the authors write. The study's findings support the hypothesis that invertebrate pattern-recognition capabilities are more flexible than previously assumed.
They also document evidence for horizontal gene transfer (HGT) between the hosts and the symbiotic dinoflagellates. Given that the associations between the two organisms have evolved over millions of years, HGT is extremely likely to have occurred. The researchers found 275 HGT candidate genes specific to Aiptasia, plus another 548 candidates believed to have been transferred from a nonmetazoan source to a basal cnidarian in the evolutionary past and shared among a large number of cnidarians.
The researchers conclude that the variety of conserved features found in their analysis should help to illuminate the evolution of many kinds of symbiotic anthozoans, not just Aiptasia-Symbiodinium holobionts, thus enhancing knowledge of reef systems, among the most important marine ecosystems.
More information: "The genome of Aiptasia, a sea anemone model for coral symbiosis." PNAS 2015 ; published ahead of print August 31, 2015, DOI: 10.1073/pnas.1513318112
Abstract
The most diverse marine ecosystems, coral reefs, depend upon a functional symbiosis between a cnidarian animal host (the coral) and intracellular photosynthetic dinoflagellate algae. The molecular and cellular mechanisms underlying this endosymbiosis are not well understood, in part because of the difficulties of experimental work with corals. The small sea anemone Aiptasia provides a tractable laboratory model for investigating these mechanisms. Here we report on the assembly and analysis of the Aiptasia genome, which will provide a foundation for future studies and has revealed several features that may be key to understanding the evolution and function of the endosymbiosis. These features include genomic rearrangements and taxonomically restricted genes that may be functionally related to the symbiosis, aspects of host dependence on alga-derived nutrients, a novel and expanded cnidarian-specific family of putative pattern-recognition receptors that might be involved in the animal–algal interactions, and extensive lineage-specific horizontal gene transfer. Extensive integration of genes of prokaryotic origin, including genes for antimicrobial peptides, presumably reflects an intimate association of the animal–algal pair also with its prokaryotic microbiome.
Journal information: Proceedings of the National Academy of Sciences
© 2015 Phys.org