Single-cell technologies advance the value of genomics
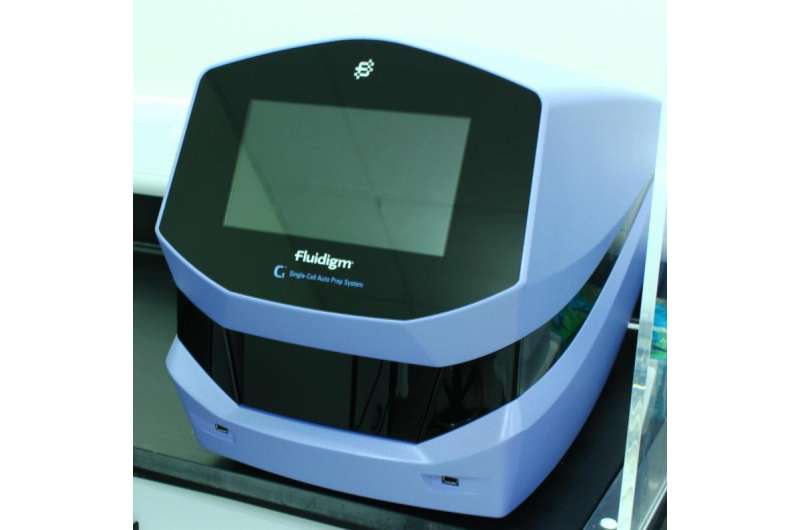

Biologists are looking to extract as much information as possible from small amounts of valuable biological material, and to understand biological responses at higher levels of resolution. The Genome Analysis Centre has been working to reduce the input requirements for DNA and RNA sequencing projects down to the single-cell level by introducing the Fluidigm C1 single-cell system, FACs-in-a-petri CellSorter and the Labcyte Echo microscopic liquid handler.
The new technologies enable the advancement of nucleic acid research methods and open up new areas of biological enquiry for the Institute and wider science community.
Providing researchers with access to better genome assemblies (commonly limited by poor input DNA), and through low input and single-cell transcriptomics, to an unprecedented understanding of the biochemical changes within cells and how to model this. The low input and single-cell transcriptomic strategy has great potential in aiding multidisciplinary approaches e.g. cell biology, genetics, genomics, biochemistry, virology, immunology, systems biology and eventually synthetic biology.
Working with low input amounts of DNA and RNA require specialist equipment, technical expertise, and precise quality control (QC) - especially to prevent and identify contamination. The selection of specific single cells is itself technically challenging, especially for the wide variety of species samples that TGAC (ww.tgac.ac.uk) deals with. In response, the Fluidigm C1 will facilitate demand through its unique automated microfluidic approach to single-cell research.
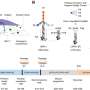
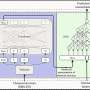
The Fluidigm C1 platform is a self-contained automated system that sorts a mixture of cells by size and creates sequencing libraries. It has a proven track record in vertebrate biology, and TGAC will use it to investigate at single cell resolution how the vertebrate immune system responds to antigens in order to improve vaccination strategies for farm animals.
The FACS-in-a-petri system provides flexibility across the range of projects and organisms on which TGAC works to select specific cells from a mixture outside the parameters possible with the C1. The Labcyte Echo 525 platform is able to dispense nanolitre volumes of reagents allowing the generation of libraries in small volumes which minimise costs while maximise yield.

These new resources will underpin a range of research projects harnessing the power of genomics, generating impact widely though many diverse user groups. Enhancements to TGAC's National Capability in Genomics supports the institute's goal to enable researchers tackling the global challenges of healthy ageing, food and energy security, sustainability and environmental change.
By offering widened access to the equipment in a multi-disciplinary approach, TGAC will not only further their own research impact, but also further enhance the capability and research incentives of other institutes in Norwich Research Park.
Tarang Mehta, Project Lead and Postdoctoral Research Scientist in the Vertebrate & Health Genomics Group at TGAC, said: "It is fantastic that we have acquired such systems which will not only advance 'omics' technology development, but also serve as a valuable resource for studying key research questions associated with animal, human, and plant health and disease."
Matthew Clark, Plant & Microbial Genomics Group Leader at TGAC, said: "These technologies open up the opportunity to understand biology at increasingly high resolutions. My group is looking forward to using them to dissect plant immune transcriptional responses to biotic and abiotic stresses."
TGAC have also looked to enhance methodologies for genome sequencing and assembly of large genomes. Specialist DNA extraction protocols for the generation of long fragments can be expensive, difficult, require specific equipment and involve the use of cryogenics and hazardous chemicals. These requirements and the lack of trained personnel can lead to compromises on quality and delayed projects.
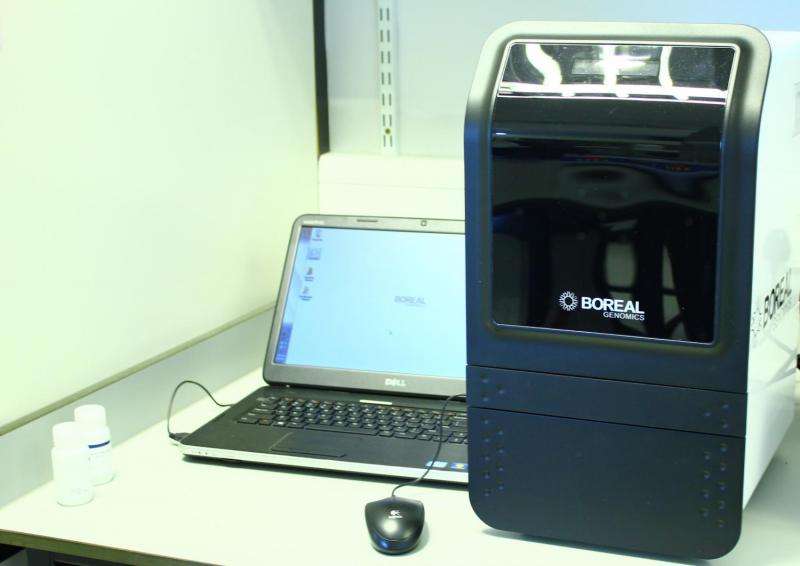

The Boreal Genomics Aurora will assist with DNA extraction required for the preparation of high quality, high molecular weight DNA libraries. It can rapidly size DNAs as large as 60kb and size select DNA molecules over 100kb to separate them from smaller fragments and inhibitory compounds.
The integration of systems can facilitate high-throughput DNA extraction through fast quantification, QC and precise size selection. For challenging samples, TGAC has also added alongside their suite of NGS technologies, a 3D scanner and printer combination to scan existing plasticware e.g. sample plates, magnet holders etc. and rapidly prototype new modifications that match TGAC's needs in DNA extraction.
Daniel Swan, Head of Platforms & Pipelines at TGAC, said: "The continued investment in cutting-edge platforms at TGAC means we are uniquely placed to address the challenges associated with complex genomics projects and we look forward to opening the platforms to the wider research community."
TGAC is strategically funded by BBSRC and operates a National Capability to promote the application of genomics and bioinformatics to advance bioscience research and innovation.
More information: www.fluidigm.com/products/c1-system
Provided by The Genome Analysis Centre