Two-armed control of ATR, a master regulator of the DNA damage checkpoint

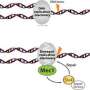
Our genomes encounter DNA damaging events continuously. As damage accumulates, our cells react and stop the cell division process, giving repair mechanisms time to act. To achieve this, two master protein kinases called ATR and ATM, encoded by well-known tumor suppressor genes, activate a cascade of phosphorylation events. The downstream readout of these checkpoint cascades is well characterized, yet the regulation of the master kinase itself has remained enigmatic. Nicole Hustedt, a student in the laboratory of Susan Gasser, examined this question with a combination of sophisticated yeast genetics and phosphoproteomics. She found that the key regulator of the yeast Mec1 kinase, the yeast ATR homologue, is a phosphatase that directly reverses the modification of many damage-induced Mec1 kinase targets. The identified phosphatase, called PP4, not only dephosphorylates Mec1 targets, but directly interacts with the kinase, fine-tuning its activity during the cell cycle.
This suggests a remarkable regulatory principle: the master kinase ATR/Mec1 appears to be counterbalanced by a phosphatase, with which it forms a complex. This chimera fine-tunes the DNA damage checkpoint response to allow cells to survive DNA damage.
Appropriate duplication of the genome is an absolutely critical process for the survival of an organism. However, all too easily errors and lesions in our chromosomes accumulate, leading to cell death, or loss of identity and cancer. To ensure the integrity of the genome, checkpoint kinases control the cell division cycle, through a cascade of phosphorylation events. These delay cell division and stimulate appropriate and timely repair.
In order to uncover the mechanisms that regulate the upstream DNA damage checkpoint kinase, Susan Gasser and her group at the Friedrich Miescher Institute for Biomedical Research took advantage of a powerful genetic technology called E-MAP that has been optimized in budding yeast. Combining synthetic genetic interactions with phosphoproteomics, carried out by FMI's Protein analysis facility, a novel regulatory interaction that controls the DNA damage checkpoint during the replication of the genome was discovered.
The large-scale combinatorial genetic approach called E-MAP allows one to identify pairs of genes that act together or antagonistically in a given process. This was applied to the Mec1-regulated DNA damage response. The FMI graduate student, Nicole Hustedt, found that only one single pair of genes could efficiently counteract defects in the Mec1-mediated checkpoint cascade, when it was activated by replication fork damage. The two genes detected were shown to encode subunits of PP4, a phosphatase. The study, now published in Molecular Cell, showed that the PP4 phosphatase, Pph3-Psy2, regulates a large number of the phosphorylation targets of Mec1. It thus contributes to a balance between phosphorylation and de-phosphorylation. This allows a rapid and effective response to damage induced checkpoint activation and, eventually, resumption of the cell cycle. Intriguingly, they also found that the PP4 phosphatase physically interacts with Mec1, forming a complex that contains both a kinase and a phosphatase.
"The two enzymes act in a coordinated yet opposing manner, which easily allows for a fine tuning of Mec1 activity during the cell cycle," comments Gasser. "The counteracting modifications are the yin and the yang of the DNA damage checkpoint."
The mechanisms controlling the cell cycle and activating checkpoints are conserved between organisms. Mec1 and Pph3-Psy2 have mammalian homologues that also interact. "By learning about the interactions in yeast, we could make the leap to human cells, and show that the regulation of the mammalian Mec1 homologue, ATR, uses the same mechanism," comments Gasser. "This helps us to understand the processes that ensure the integrity of the genome, preventing oncogenic transformation."
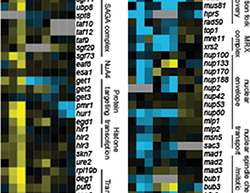
Epistatic miniarray profile – E-MAP
In an E-MAP screen scientists are looking for mutant proteins that either alleviate or worsen the effect of another, known mutation. Such genetic interactions are called epistatic or additive, respectively, and their analysis allows insights into the structure and function of pathways. E-MAP measures genetic interactions of proteins in a high-throughput and semi-quantitative fashion. The basis of the screen is a quantitative phenotype, e.g. growth. The screen then identifies mutant combinations that fare better or worse under certain circumstances. The data is then clustered hierarchically, thus allowing the identification of genes that directly or less directly interact with the first mutation. In the above mentioned screen, the scientists measured growth of mutants in the absence of functional Mec1. The screen thus identified proteins masking the effects of the Mec1 mutation.
More information: Hustedt N, Seeber A, Sack R, Tsai-Pflugfelder M, Bhullar B, Vlaming H, van Leeuwen F, Guénolé A, van Attikum H, Srivas R, Ideker T, Shimada K, Gasser SM (2014) "Yeast PP4 interacts with ATR homologue dc2-Mec1 and regulates checkpoint signaling." Mol Cell DOI: 10.1016/j.molcel.2014.11.016
Journal information: Molecular Cell