What's for dinner? Rapidly identifying undescribed species in a commercial fungi packet

For lovers of wild foods, autumn harks a season of bounty. Fungi of dizzying variety erupt from wood and soil, luring intrepid collectors to woodlands in search of elusive but delectable wild mushrooms. Part of their appeal lies in the allure of the treasure hunt, and their mysterious not-quite-meat, not-quite-vegetable qualities that belie an almost otherworldly existence. But are the mushrooms which you are eating known to science?
The Fungi Kingdom is enormously diverse yet vastly underdocumented – although some estimates range up to 10 million species, only about 100,000 species have been described. Mushrooms are one the most conspicuous and well known groups of Fungi and make up around 16,000 named species, but only a handful of these species are well documented. With estimated rates of Fungi extinction exceeding current rates of description, the enormity and urgency of the task of accurate identification cannot be overstated. New approaches that accelerate the documentation and description of new species are desperately needed before it is too late.
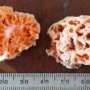
So how well do we really know the mushrooms? In an article published today in PeerJ, Mycologists Bryn Dentinger and Laura Martinez-Suz from the Royal Botanic Gardens, Kew in London set out to ask this question by using DNA-based taxonomy. But instead of venturing into the wild in search of never before seen species, they simply went to a local grocer and bought a packet of dried porcini whose intended destiny was more likely a rich risotto than a DNA sequencer.
Some of the most sought-after of wild mushrooms are the sweet and nutty Boletus edulis and allies, often referred to by the Italian common name porcini. Dentinger has been studying porcini for over 10 years and knew that if the porcini in this packet originated in China, they were likely to be made up of unnamed species. But how many species could be detected in the packet and how quickly could diagnosing and describing them be accomplished?

Drs Dentinger and Suz arbitrarily selected 15 pieces of mushroom from the packet and sequenced the fungal DNA barcode region for each. They then compared these sequences to sequences in the International Nucleotide Sequence Database and classified them based on evolutionary relationships. This revealed three distinct species, none of which were known to science, or had scientific names. To expedite the formal naming process required by the International Code of Nomenclature for algae, fungi and plants, the researchers used a rapid e-publishing tool that facilitates immediate name registration and species diagnosis to satisfy the rules of the Code.
"This study demonstrates that the whole procedure, from unknown mushrooms to names, can be done rapidly – we did this in under a week, but if push came to shove, it could be done in a day" said Dr Dentinger.

Hundreds of thousands of tons of porcini are collected from the wild and sold around the world every year, most of it ending up in Europe and North America. Although typically composed of five well-known species native to these regions (B. aereus, B. edulis, B. pinophilus, B. reticulatus, B. rex-veris), around half of all porcini traded in Europe originates in China. These porcini, typically collected from the wild in Yunnan province, have been exported to Europe since the 1970s yet until last December, none of them had scientific names. The researchers named them with Chinese epithets referring to local common names for porcini (Boletus meiweiniuganjun, Boletus bainiugan) and the Chinese word for 'edible' (Boletus shiyong).
As Dr Dentinger put it, "our results demonstrate just how ubiquitous unknown fungal diversity is – it can literally be found right under our noses."
The researchers hope that by demonstrating the rapid identification and naming of new Fungi species in this way, others in the community will be inspired to continue the important work of identifying new Fungi species before they disappear.
More information: Dentinger and Suz (2014), What's for dinner? Undescribed species of porcini in a commercial packet. PeerJ 2:e570; DOI: 10.7717/peerj.570
Journal information: PeerJ
Provided by PeerJ