Subseafloor bacteria survive by over-activating DNA-repair and antibiotic target genes
The subseafloor is home to over 1/3 of the bacteria on the planet, but up until recently it was unclear if this huge microbial biosphere was alive and dividing. Now the same group that demonstrated this activity has shown that bacteria from the hostile sea-floor environment have adapted by over-activating stress response and DNA-repair mechanisms, to cope with the harsh conditions.
Subseafloor sediment contains the Earth's largest habitat for microbial life – over 1/3 of all the planets microbial biomass. By drilling deep into the sea floor and taking samples, it can be proven that the subseafloor contains a variety of microbial lifeforms, but it's only in the last year that researchers have proven that sea floor microbes are actually active in in their natural sea-bed situation - it is difficult to analyse lifeforms which live hundreds of metres below the sea surface because of their low activity levels. A group of researchers at Woods Hole Oceanographic Institute and University of Delaware* developed techniques to analyse the messenger
RNA (mRNA) molecules produced by subseafloor microbes. Unlike DNA, which is a fairly robust molecule that can survive intact for thousands of years under certain conditions, mRNA (messenger RNA) has a short half-life. It is produced by cells, which are "turning on" genes, so it is an indication that genes are active. This means that mRNA can be used as evidence (a proxy) for present biological activity.
Lead researcher William Orsi said: "This is the largest microbial biosphere on Earth, composed of cells living deep beneath the surface. We have recently shown for the first time that these cells, the "deep biosphere", are actually dividing and not in a dormant state. This means that the deep biosphere is active and due to its sheer size likely plays an important role in global elemental cycles over geological timescales". Now in a presentation to the Goldschmidt conference in Sacramento, California, Dr Orsi will detail just how these seabed bacteria had managed to survive in such an inhospitable environment.
"It's a really difficult environment to study, so understanding how microbes survive there has been a puzzle" he said, "but we have discovered that they ramp-up some coping mechanisms which have helped them adapt to this stressful environment, where they exist under high pressure and are starved of nutrients".
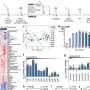
The group sampled drill cores from the continental shelf off the coast of Peru. They compared gene expression at several depths spanning 5-159 meters below the seafloor. They found that the expression of DNA repair genes, such as recA, increases with the amount of time the microbes have been buried in the seafloor.
Dr Orsi continued: "Subseafloor microbes have adapted to live in especially harsh conditions. We found that they significantly overexpress genes involved in cellular stress responses like recA. This gene is a central to the bacterial "SOS response", which is a way bacteria cope with many different environmental stressors including antibiotics. We have found that subseafloor microbes increasingly express this gene with time after they become "buried alive" in the subseafloor".
"High-throughput omics techniques are proving to have a range of applications in sedimentary systems. For example, marine sedimentary paleogenomics is a new field, which is opening windows into the past effects of climate on marine life. Dr Marco Coolen is the leader of this new field, and we worked together to analyse ancient plankton DNA from the Black Sea. We showed that it opened to the oceans around 9,600 years ago and that historical large-scale climate changes had a significant effect on marine plankton. These techniques are difficult, but they can tell us how biology responds to climate changes over geological timescales. Looking into the past can help us predict the future effects of climate change on marine life".
Provided by European Association of Geochemistry