April 18, 2014 report
Researchers create methylation maps of Neanderthals and Denisovans, compare them to modern humans
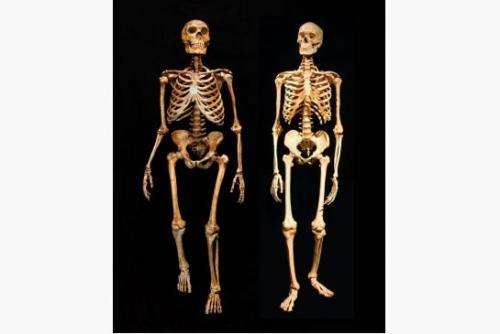
(Phys.org) —A team of Israeli, Spanish and German researchers has for the first time created a map of gene expression in Neanderthals and Denisovans and has compared them with modern humans. In their paper published in the journal Science, the team describes how they applied epigentics to the study of our two closest known ancestors and discovered variations that might account for their differences in body shape and susceptibility to some modern neurological diseases.
Scientists know that it's not just our DNA structure that determines how we look and what we're capable of doing, there's another factor involved—the expression of our genes—they can be turned on or off at some point, allowing or preventing certain traits from developing. This process is known as DNA methylation—where methyl group chemicals attach to DNA and prevent them from behaving as they would otherwise. In this new effort, the researchers looked at methylation in Neanderthals and Denisovans to learn more about how they might have been different from us.
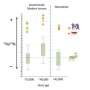
Studying methylation in preserved fossils involves noting the way the methyl chemical cytosine decays over long periods of time. Unmethylated cytosines decay to one type of chemical while unmethylated cyctones decay to another. By measuring the amounts of the two resultant chemicals found in fossilized bone fragments, the researchers were able to create methylation maps of Neanderthals and Denisovans, which they then compared with similar maps for modern humans.
The comparisons revealed differences in approximately 2000 different regions, though one in particular stood out—an HoxD cluster that prior research has shown plays an important role in the development of body structure—a finding that could help explain the shorter, stouter limbs (and other features) of our extinct cousins. Interestingly, the team also found that some of the highly methylated regional areas that appear in modern humans do not appear in either Neanderthals or Denisovans, regions that have been associated with neurological disorders such as schizophrenia and autism—a finding that may help shed some light on their source.
Unfortunately, the maps created by the research team are still incomplete, they only had a few bone fragments to work with—they're hoping future studies (and fossil finds) will reveal more. In the meantime, they plan to conduct similar work on other species, such as horses, to help reveal the types of methylation that occurred as they were domesticated.
More information: Reconstructing the DNA Methylation Maps of the Neandertal and the Denisovan, Science, DOI: 10.1126/science.1250368
ABSTRACT
Ancient DNA sequencing has recently provided high-coverage archaic human genomes. However, the evolution of epigenetic regulation along the human lineage remains largely unexplored. We reconstructed the full DNA methylation maps of the Neandertal and the Denisovan by harnessing the natural degradation processes of methylated and unmethylated cytosines. Comparing these ancient methylation maps to those of present-day humans, we identified ~2000 differentially methylated regions (DMRs). Particularly, we found substantial methylation changes in the HOXD cluster that may explain anatomical differences between archaic and present-day humans. Additionally, we found that DMRs are significantly more likely to be associated with diseases. This study provides insight into the epigenetic landscape of our closest evolutionary relatives and opens a window to explore the epigenomes of extinct species.
Journal information: Science
© 2014 Phys.org