Microsatellites are repetitive, but the lab work doesn't have to be
Microsatellites are molecular markers with numerous applications in biological research. In studies of both plants and animals, they can be used to investigate speciation, gene flow among populations, mating systems, and parentage, as well as many other questions. A new protocol created by researchers at the University of Cincinnati and several other institutions improves the efficiency of current methods, allowing quicker and cheaper development of microsatellite markers for any species of interest.
Microsatellites, which consist of repeating units of two to six base pairs of DNA, are desirable molecular markers because they are highly variable and co-dominant, allowing researchers to determine both parental contributions to an individual's genotype. However, microsatellite markers are generally species-specific and frequently must be developed for each study from scratch, which can be a challenging task. Additionally, once microsatellite markers have been developed, the associated lab work is often time-consuming and expensive.
University of Cincinnati professor Theresa Culley and colleagues have established a method that increases the ease and affordability of microsatellite marker development by reducing the number of steps involved in the overall process. The new protocol is described in the October issue of Applications in Plant Sciences.
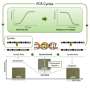
Previously, researchers would generally design primers for specific microsatellite markers and label each primer pair with fluorescent dyes in separate PCR reactions (for each individual included in the study). In studies including approximately 10 microsatellite markers and several hundred individuals, this would necessitate up to several thousand PCR reactions. The fluorescently labeled PCR products (i.e., amplified microsatellites) for each individual would then be combined, or pool-plexed, for genotyping.
In the new protocol, multiple microsatellites can be fluorescently labeled and amplified in the same PCR reaction, significantly reducing the total number of PCR reactions required for each study.
"The main advantage of this protocol," says Culley, lead author of the study, "is really in streamlining the primer testing process and making it easier and more economical for researchers with limited funding to do microsat studies."
Culley adds that the protocol is also advantageous because it permits considerable flexibility and experimentation during the primer testing process. Says Culley, "The protocol is significant because it offers a way that researchers can easily switch the fluorescent tags on their primers during testing with minimal cost, as opposed to spending $70 - $80 per primer of a specific dye color and then having to reorder that primer again with a different color."
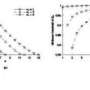
In the study, the authors also examine the effect of different thermocycler conditions (namely, cycle number and annealing temperature) on PCR success, in order to identify the conditions that optimize the PCR process.
According to Culley, the new method can also be applied to the development of additional types of molecular markers.
"Although it was developed for microsatellite markers, it can conceivably be used for any other type of marker as well," notes Culley. "For example, in our lab we have been using it to tag primers for the self-incompatibility locus in various plant species."
This method complements other recently developed techniques for microsatellite research, all of which serve to streamline the primer development process.
More information: Theresa M. Culley, Trevor I. Stamper, Richard L. Stokes, Jessica R. Brzyski, Nicole A. Hardiman, Matthew R. Klooster, and Benjamin J. Merritt. 2013. An efficient technique for primer development and application that integrates fluorescent labeling and multiplex PCR. Applications in Plant Sciences 1(10): 1300027. DOI: 10.3732/apps.1300027
Journal information: Applications in Plant Sciences
Provided by American Journal of Botany