Protein signaling between soybean root hairs, bacteria reveals core cellular processes

(Phys.org)—Understanding what happens to a soybean root hair system infected by symbiotic, nitrogen-fixing soil bacteria, Bradyrhizobium japonicum, could go a long way toward using this symbiosis to redesign plants and improve crop yields, benefitting both food and biofuel production. Because of their extensive genomes, it is especially difficult to use conventional proteomic technologies to get meaningful information from plants. With the availability of a complete soybean genome, soybean root hairs represent an excellent model for the study of single-cell systems biology.
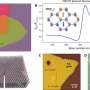
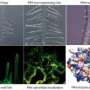
Legume root hairs primarily are involved in water and nutrient uptake from the soil but also are the dominant infection site of symbiotic rhizobia. This infected area forms a novel organ—the nodule—where bacteria fix nitrogen for the host, acting as built-in fertilizer. At EMSL, scientists, as part of an onging collaboration with the Stacey Laboratory, employed the ultra-sensitive liquid chromatography-Fourier transform mass spectroscopy platform to characterize the soybean root hair proteome and determine root hair cellular signaling cascade responses to rhizobial colonization and infection. Stripped roots (with no root hairs), non-inoculated soybean root hairs, and inoculated root hairs (with B. japonicum) were watched for changes over a 72-hour period.
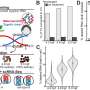
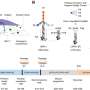
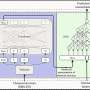
Nine time points were analyzed separately then combined to establish the root hair reference proteome map. The process cataloged more than 5700 proteins, some specific to the root hair, involved in cell functionality, such as nutrient uptake. The scientists also used EMSL's ultra-sensitive phosphoproteomic platform coupled with eight-plex iTRAQ methodology, which labels all peptides in up to eight different biological samples, to track how the cell's defense system was suppressed to allow nodules to form on roots. Signals between root hair cells and bacteria orchestrate complex and rapid cellular changes. Thus, these studies are important for piecing together the necessary genetic and protein targets that compose this beneficial symbiotic relationship.
More information: Brechenmacher L, et al. Identification of soybean proteins from a single cell type: The root hair. Proteomics 12(22):3365–3373, (2012). DOI: 10.1002/pmic.201200160 3365
Nguyen T. et al. Quantitative phosphoproteomic analysis of soybean root hairs inoculated with Bradyrhizobium japonicum. Molecular and Cellular Proteomics 11(11):1140-1155, (2012). DOI: 10.1074/mcp.M112.018028
Journal information: Proteomics , Molecular and Cellular Proteomics
Provided by Environmental Molecular Sciences Laboratory