May 17, 2012 report
Group finds circadian clock common to almost all life forms
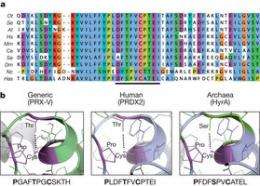
(Phys.org) -- A group of biology researchers, led by Akhilesh Reddy from Cambridge University have found an enzyme that they believe serves as a circadian clock that operates in virtually all forms of life. In a paper published in the journal Nature, they describe a class of enzymes known as peroxiredoxins which are present in almost all plants and other organisms and which appear to serve as a basic ingredient in non-feedback loop biological clocks.
Up till now, researchers have not been able to find any kind of common biorhythmic clock among the Earth’s varied organisms, each class seemed to have its own. They did find though that one common feature of most was a feedback loop, which is where genes are transcribed before being translated into proteins which then build up until they reach a tipping point. Once that happens, transcription is turned off and the enzyme goes dormant. This cycle, for most organisms occurs on a twenty four hour basis, and is responsible for such things as the feelings of sleepiness or hunger in people that occur at roughly the same time each day.
But now, this new research suggests that the true clock controlling behavior in virtually every imaginable plant, animal, fungus, etc. has its roots in an enzyme whose purpose is to help clean up residue left over from the ravages of antioxidants.
Peroxiredoxins, which exist in virtually all life forms, are enzymes that cycle between two states depending on whether they have reacted recently with hydrogen peroxide, or not. The researchers found that this cycle occurs on a roughly twenty four hour cycle in all of the organisms they’ve tested to date. What’s more, the cycle continued even in the absence of light, proving that it’s not part of a feedback loop. Unfortunately, the team has not yet been able to show how or if the enzyme controls other clock mechanisms that are a part of feedback loops.
The team suggests that peroxiredoxins developed their cyclical behavior just after organisms began to develop some two and half billion years ago that were able to handle the increased amounts of oxygen that had begun to appear in the atmosphere in a time period known as the Great Oxidation Event; the time when bacteria developed photosynthesis and began pumping out oxygen. Those organisms that managed to survive had to develop a means of dealing with the damage caused by antioxidants, and thus was born the role of peroxiredoxins. And because oxygen levels rose and fell on a regular daily schedule, the enzymes developed a clock over time to help predict when to go to work, and when to remain dormant, thus paving the way for the first circadian clock.
More information: Peroxiredoxins are conserved markers of circadian rhythms, Nature (2012) doi:10.1038/nature11088
Abstract
Cellular life emerged ~3.7 billion years ago. With scant exception, terrestrial organisms have evolved under predictable daily cycles owing to the Earth’s rotation. The advantage conferred on organisms that anticipate such environmental cycles has driven the evolution of endogenous circadian rhythms that tune internal physiology to external conditions. The molecular phylogeny of mechanisms driving these rhythms has been difficult to dissect because identified clock genes and proteins are not conserved across the domains of life: Bacteria, Archaea and Eukaryota. Here we show that oxidation–reduction cycles of peroxiredoxin proteins constitute a universal marker for circadian rhythms in all domains of life, by characterizing their oscillations in a variety of model organisms. Furthermore, we explore the interconnectivity between these metabolic cycles and transcription–translation feedback loops of the clockwork in each system. Our results suggest an intimate co-evolution of cellular timekeeping with redox homeostatic mechanisms after the Great Oxidation Event ~2.5 billion years ago.
Journal information: Nature
© 2012 Phys.Org