Using 'proton grease' to spin-up a molecular rotor
When designing the tiniest of possible machines, scientists have had far more success in creating molecular-size brakes than accelerators. But a team at the University of South Carolina has figured out how to really hit the gas pedal.
Reporting in the Journal of the American Chemical Society, they use what they call 'proton grease' to make a molecular rotor spin faster – by a factor of ten million. The advance is a significant signpost on the road toward functional synthetic molecular machines, said Ken Shimizu, lead author and professor in the department of chemistry and biochemistry in the College of Arts and Sciences at USC.
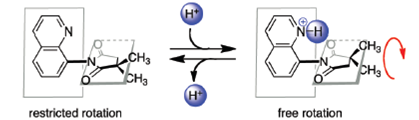
The team constructed their rotor by combining quinoline and succinimide subunits. At first blush it might look like rotation in the resulting molecule would be essentially unrestricted, but the appearance is deceiving. The partial charge on the carbonyls has highly unfavorable overlap with the quinoline nitrogen when the molecule is in a planar transition state.
But when the nitrogen is protonated, the planar rotamer is stabilized – leading to a huge increase in the rotational rate.
"We designed it to have more favorable overlap after protonation, so we expected it to speed up," said Shimizu. "But we never anticipated the magnitude of the increase – we were surprised by how well it worked, which is a rare thing to say."
The barrier to rotation dropped from about 22 kcal/mol to 13 kcal/mol upon titration with acid. The increase in rotation speed was so dramatic, covering seven orders of magnitude, that the team had to cobble together two different measurement techniques – one for fast rotation, and another for slow.
And as they show in their research article, the speed-up is reversible: addition of base restored the barrier to rotation.
The field of molecular devices is in its infancy, but the possibilities are tantalizing. "We're kind of making cogs right now, but we're moving toward being able to make more complex devices," said Shimizu. "People are imagining all sorts of electronic and mechanical devices based on single molecules, and we're building up this toolbox to be able to create machines and devices on the molecular scale."
Despite coming up with the evocative term 'proton grease,' Shimizu recognizes the name carries potential baggage as well. "It makes it more accessible to use these analogies about brakes and grease and things like that – but it's a little bit dangerous too," he said.
"When people talk about in these molecular machines, they often assume that you can take an engine, for example, and just scale it down to a molecular size. And conceptually you can, but the actual physics behind that falls apart," said Shimizu. "We don't have friction at that scale, for example."
"But that's actually the most exciting part, too," he added. "When you start making things on that scale, the rules are completely different."
"By controlling motion on the molecular level, people imagine that you could make switches, memory, transformers – all sorts of electronic and mechanical devices as single molecules."
More information: Proton Grease: An Acid Accelerated Molecular Rotor, J. Am. Chem. Soc., 2012, 134 (8), pp 3675–3678. DOI: 10.1021/ja2120184
Abstract
A molecular rotor was designed that rotates 7 orders of magnitude faster upon protonation. The quinoline rotor is based on a rigid N-arylimide framework that displays restricted rotation due to steric interaction between the quinoline nitrogen and imide carbonyls. At rt (23 °C), the rotor rotates slowly (t1/2 = 26 min, ΔG‡ = 22.2 kcal/mol). However, upon addition of 3.5 equiv of acid the rotor rotates rapidly (t1/2 = 2.0 × 10–4 s, ΔG‡ = 12.9 kcal/mol). Mechanistic studies show that this dramatic acid catalyzed change is due to stabilization of the planar transition state by the formation of an intramolecular hydrogen bond between the protonated quinoline nitrogen (N+—H) and an imide carbonyl (O═C). The acid catalyzed acceleration is reversible and can be stopped by addition of base.
Provided by University of South Carolina