New paper examines poison resistance in snakes around the world

(PhysOrg.com) -- A new study by University of Notre Dame biologist Michael Pfrender and a team of researchers from the University of Nevada, Reno; Utah State University; and the University of Virginia suggests that snakes from different regions of the world have evolved a similar, remarkable resistance to a deadly neurotoxin.
The finding, which appeared in the Proceedings of the National Academy of Sciences, greatly increases scientists’ understanding of the genetic basis of adaptation and is a model for understanding the limits to adaptation and the degree to which evolutionary responses are predictable.
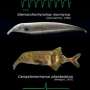
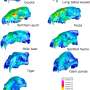
Pfrender and colleagues found species of snakes in North, Central and South Americas and Asia that are able to feed on amphibians that secrete a deadly neurotoxic poison, tetrodotoxin or TTX. These snakes have similar mutations in a key sodium-channel gene that makes them highly resistant to TTX. These mutations prevent TTX from blocking the sodium channels in muscle, which would otherwise immobilize the snakes by paralyzing nervous and muscle tissue.

“The key finding is that adaptive evolution is constrained by the functional properties of the genes involved in these evolutionary responses,” Pfrender said. “While there are many possible mutations that can improve fitness, in this case resistance to the neurotoxin TTX, many of these mutations have a cost because they change the normal function of the genes. So, when we look at multiple species that have independently adapted to TTX, we see a very similar, and limited, set of mutations involved. The story is one of repeated evolutionary change that occurs through a limited set of changes at the molecular level.”
The study stems from Pfrender’s interest in understanding how organisms deal with environmental change through adaptive evolution.
“We would like to know what the underlying genetic mechanisms are, and what the limits are to these adaptive responses,” he said. “Ultimately, we would like to develop a predictive framework to gauge when natural populations will be able to evolve rapidly enough to persist in a changing environment and when the environmental change is too fast or too strong, leading to local extinction.”
An understanding of how organisms deal with environmental change is relevant to the major themes of Notre Dame’s Environmental Change Initiative and to the Eck Institute for Global Health, which examines disease resistance coupled with human health.
“Many organisms are exposed to toxic chemicals in their environment, and this system is a model for understanding how they cope with this challenge through evolutionary change,” Pfrender said. “A good example of the application of this knowledge is when we are trying to understand how parasites acquire drug resistance. How do they do it and what are the limits to this response? Can we create more effective drug strategies that capitalize on these functional constraints, making it more difficult for parasites to evolve resistance?”
Pfrender and the Utah State researchers plan to study more snake species and to expand their research to a number of other species, including insects that prey on the toxic eggs of salamanders. They also are examining other genes closely related to the sodium channel genes that are the focus of the PNAS study to expand their understanding of how adaptation occurs.
Provided by University of Notre Dame









-1.jpg)
.jpg)