Making nature's best better to produce biofuels
If a tree falls in the forest and there are no enzymes to digest it, does it break down?
It's a question that has important ramifications for the renewable energy industry. Engineers are studying methods to transform non-food plant material into transportation fuel. Think alfalfa stalks or wood-chips (which have energy contained in a molecule humans can't digest called cellulose), as opposed to the edible corn grains that are used in the production of ethanol for biofuels.
"Cellulose in the biosphere can last for years," said Gregg Beckham, a scientist in the National Bioenergy Center at the U.S. Department of Energy's National Renewable Energy Laboratory (NREL). "It's really tough, and we want to know why at the molecular scale."
Despite the strength of plant cell walls made of this tough molecule cellulose, over eons, fungi and bacteria have evolved enzymes to convert abundant cellulosic plant matter into sugars to use as an energy source to sustain life.
Breaking down in the lab
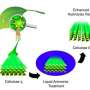
Unfortunately, these particular enzymes don't work fast enough to break down cellulose at a pace (and price) that is competitive with fossil fuels ... yet. So, computational scientists at NREL set about trying to understand and create enhanced, "designer" enzymes to speed up biofuel production and lower the cost of biomass-derived fuel to serve the global population.
"It's a Goldilocks problem," Beckham said. "The enzymes have to be 'just right,' and we're trying to find out what 'just right' is, why, and how to make mutations to the enzymes to make them most efficient."
Supercomputed proteins
In a series of linked projects, researchers used the National Science Foundation-supported Ranger supercomputer at the Texas Advanced Computing Center and Energy Laboratory's Red Mesa system to simulate the world of enzymes. The researchers explored enzymes from the prodigiously plant-digesting fungus Trichoderma reesei and the cellulose-eating bacteria Clostridium thermocellum. Both of these organisms are effective at converting biomass to energy, though they use different strategies.
"Nature cleverly designed machinery for single-cell organisms to locate cellulose, then secrete large enzyme complexes that hold the cells near biomass while the enzymes degrade it," Beckham said.
The bacteria forms scaffolds for its enzymes, which work together to break apart the plant. The fungal enzymes, on the other hand, are not tethered to a large complex, but act independently.
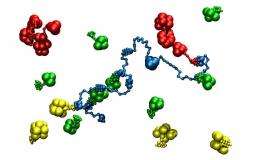
It isn't clear how the enzyme scaffolds form, so the researchers created a computational model of the active molecules and set them into motion in a virtual environment. Contrary to expectations, the larger, slower-moving enzymes lingered near the scaffold longer, allowing them to bind to the frame more frequently; the smaller ones moved faster and more freely through the solution, but bound less often.
The results of the study, led by NREL researchers Yannick Bomble and Mike Crowley, were reported in the Journal of Biological Chemistry in February 2011. The insights are being used in the creation of designer enzymes to make biomass conversion faster, more efficient and less expensive.
Unexplored enzyme function
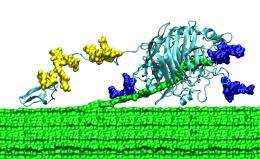
The scientists also studied parts of the enzyme called the carbohydrate binding molecule--a sticky "foot" that helps the enzymes find and guide the cellulose into their active site--and the linker region, which joins the foot to the main body of the enzyme. The carbohydrate binding molecule and linker region were long thought to play a minor role in enzyme function; yet without them, the enzyme can't convert cellulose to glucose effectively. The researchers wondered why that is.
Using the Ranger supercomputer, the researchers made several important discoveries. First, they found that the cellulose surface has energy wells that are set 1 nanometer apart, a perfect fit for the binding module. They also found that the linker region, previously believed to contain both stiff and flexible regions, behaves more like a highly flexible tether. Those insights would have been difficult to determine experimentally, but, now hypothesized and backed up with advanced computing simulations, they can be tested in the laboratory.
"It's a very messy problem for the experimentalists," said Crowley, a principal scientist at the Energy Laboratory and Beckham's colleague. "We're using rational design to understand how the enzyme works, and then to predict the best place to change something and test it."
The research addresses the enzymatic activity bottlenecks that prevent renewable energy from cellulose containing biomass from being competitive with fossil fuels. "If we can help industry understand and improve these processes for renewable fuel production, we'll be able to offset a significant fraction of fossil fuel use in the long term," Beckham said.
Provided by National Science Foundation