Reversing ecology reveals ancient environments
From hair color to the ancestral line of parasitic bacteria, scientists can glean a lot from genes. But imagine if genes also revealed where you lived or who you spent time with. It turns out they do, if you know where and how to look.
Stanford researchers with collaborators at Tel-Aviv University have now laid the foundation for opening such a window to the past using a technique called "reverse ecology." The technique uses genomic data to examine metabolic networks and pulls out proxies for reconstructing bacterial environments millions of years in the past. The work, published in the February issue of the Journal of Computational Biology, offers clues to the complex evolutionary interplay between organisms such as parasites and hosts.
"Based on reverse ecology, you can start with an organism-say, a certain bacterial species that you know nothing about ecologically. But by looking at its genome and metabolic network, you can recreate that past environment the organism lived in," said Elhanan Borenstein, lead author of the paper and a postdoctoral researcher in the Biology Department at Stanford. "And we've done this with hundreds of different species."
Researchers have used genomic data to study metabolic networks-the chemical reactions in metabolism that determine the physiological and biochemical properties of cells-in great detail. But Borenstein, with co-author Marcus Feldman, a professor of biology at Stanford, took this understanding a step further.
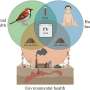
Through the metabolic network, organisms accumulate biochemical compounds from their interactions with the surrounding environment (e.g., oxygen, glutamine or sulfate). These molecules also correlate with other environmental properties like temperature and salinity. "This gives us a way to predict the biochemical environment of organisms and learn ecology from the genomic data on a large scale," Borenstein said.
The researchers collected clues about not only the organism's environment but also its relationship to other species. For example, they detected a specific signature for adaptation between a parasite and host. What's more, they could tell what kind of host the parasite was living in based on the alignment between the environment a parasite requires and that required by the host. "We can see a signal to distinguish between a mammal parasite and an insect parasite," Borenstein said. "And how the interaction evolved over time."
Now that they have a data set that reveals the current and ancient environment of hundreds of bacterial species, Borenstein and colleagues hope to use their data to discern major environmental events of the past, including key events in the history of life on Earth.
The next step is to move from looking at individual bacteria species to entire communities. In particular, Borenstein hopes to explore large collections of host-dwelling bacteria like those living in the human gut or mouth or the communities found in soils. The complicated and intricate ecology of these systems should now be accessible given the genomic data that researchers continue to unveil.
"The important thing," Borenstein said, "is that we now know there is a way to learn ecology from genomic data, and we can do this on a very large scale."
Source: Stanford University