Researchers discover way to see how a drug attaches to a cell
Sandia National Laboratories researchers John Shelnutt and Yujiang Song have discovered a better way to see where a drug attaches to a cell through a new process that produces novel hollow platinum nanostructures.
The research will appear as a paper in an upcoming issue of the German chemical journal Angewandte Chemie Int. Ed. In advance of publication, it is featured as a short "hot paper" on the journal's web site.
In their paper Shelnutt and Song describe a new way of producing porous, nanoscopic, hollow platinum spheres by using liposomes as blueprints. (Liposomes are microscopic, fluid-filled pouches that are used to deliver certain vaccines, enzymes, or drugs to the body.)
In earlier work, Shelnutt's group grew large continuous nanosheets of platinum on liposome templates, forming foam-like platinum nanostructures. This method provided no way to control shape and size.
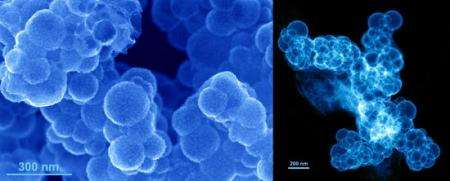
The new method reported in the paper uses a different technique to produce porous platinum nanocages with diameters up to 200 nm. Instead of large sheets, they consist of many small flat-branched platinum structures - called dendrites - which join together in a network or cage in the shape of the spherical liposome.
The liposome that Shelnutt and his team used as a blueprint consists of a double layer of lipid (detergent) molecules. The liposomes are placed in a solution containing a platinum salt. When these liposomes are irradiated with light, photocatalysts located in the narrow space between the two layers of the lipid transfer electrons to the platinum ions. The uncharged platinum atoms gather into tiny metal clumps. Once they reach a certain size, they also become active and catalyze the dendrite growth by adding more platinum atoms from the platinum salt.
Little by little, small, flat, platinum dendrites form within the double lipid layer.
"The important thing is to make sure that the number of photocatalyst molecules - and thus the number of platinum clumps - within the liposome double layer is very high," Shelnutt says. "The resulting dendrites are then close enough to join and take the shape of the liposome.
When the liposomes are broken up, the platinum spheres remain intact.
The thickness of the platinum shell around the sphere can be controlled by reducing or increasing the amount of the platinum salt placed into the solution.
Shelnutt sees many potential applications for this process, including "nanotagging" biological structures such as drug molecules.
"This would involve labeling the drug by attaching a porphyrin molecule and, after allowing the drug to bind to a cell, using light to grow a nanometer-sized particle. The nanoparticles can then be imaged with electron microscopy to reveal the location of the drug receptor molecules on the cell," Shelnutt says. "This type of nanotagging technique might be used in non-biological applications as well - such as finding flaws in semiconductor surfaces."
Source: Sandia National Laboratories





















