June 28, 2010 report
Ebola and Marburg viruses may be much older than thought
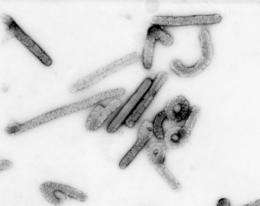
(PhysOrg.com) -- New research on the DNA of wallabies, rodents, a number of mammals and bats has found it is likely the ancestors of the Ebola and lesser-known Marburg viruses were in existence tens of millions of years ago, which is much earlier than previously thought.
The Ebola and Marburg viruses are known as "filoviruses," and result in life-threatening hemorrhaging in humans and other primates. Outbreaks occur in remote locations in Africa, and while rare they cause high fatality rates, and seem to appear out of nowhere. There are no effective treatments, and no vaccines.
It was previously thought that filoviruses were probably about 10,000 years old, with this figure based on the estimated mutation rate. The new research, by evolutionary biologist Derek Taylor and a team from the State University of New York in Buffalo, has used a different method to estimate their age.
The method is one used by paleovirologists, using remnants of virus genes found scattered within the genomes of animals. Viruses are either DNA or RNA based, and many insert their own genes into the DNA of host cells, and it had been thought that RNA-based viruses needed a gene for reverse transcriptase, an enzyme that converts their RNA to DNA in order to do this. Viruses with the gene are called retroviruses, and include the AIDS virus, HIV. Remnant viral genes from ancient retroviruses can be found in virtually every animal’s genome.
Filoviruses are RNA viruses that do not have the gene for reverse transcriptase. Taylor and co-worker Jeremy Bruenn discovered non-retrovirus genes in fungi last year, and dubbed them non-retroviral integrated RNA viruses (NIRVs). In January this year a group of researchers in Boston also reported finding RNA viruses (bornavirus) able to integrate their genes into mammalian DNA without the help of reverse transcriptase.
Taylor and his team have found “fossil” remnants of filovirus genes inside the genomes of a dozen species, but have not found any in primate species. They used genome databases to find the fossil remnants in mammals, and also confirmed their presence in a dead bat and a wallaby from a local zoo.
The team then compared the filovirus remnants in different species and found they were almost identical, which suggests the virus infected animals early in evolution, and the viral remnants were inherited by succeeding generations as the groups diverged to form separate species. For example, the house mouse and Norway rat have the same remnants in the same places in the same chromosomes, even though these diverged from each other over 12 million years ago.
Taylor said the odds of a gene inserting itself in the same place among billions of nucleotides are extremely unlikely, which means the filoviruses are much more ancient than previously thought, with the age at least in the tens of millions rather than tens of thousands.
There have been numerous studies looking for species that could harbor filoviruses without contracting the disease (known as reservoir species), and bats have been considered good candidates. The presence of NIRVs, which Taylor calls “battle scars of an infection,” could indicate the species with gene remnants could be reservoir candidates. Finding fossil remnants in New World marsupials could indicate the deaths in South America that sometimes occur after unexplained hemorrhagic fevers may be due to unidentified filoviruses.
The paper is published online in BMC Evolutionary Biology.
More information
Filoviruses are ancient and integrated into mammalian genomes, BMC Evolutionary Biology 2010, 10:193. doi:10.1186/1471-2148-10-193
© 2010 PhysOrg.com







