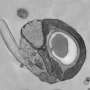
Researchers study ocean plant cell adaptation in climate change
How will plant cells that live in the oceans and serve as the basic food supply for many of the world's sea creatures react to climate change?
A University of Iowa biologist and faculty member in the Roy J. Carver Center for Comparative Genomics and his colleagues came one step closer to answering that question in a paper published in the April 9 issue of the journal Science.
Debashish Bhattacharya, professor of biological sciences in the UI College of Liberal Arts and Sciences, is studying a tiny (about one micrometer in diameter) and diverse group of organisms called picoeukaryotes. So far, he has found that organisms from two isolated groups of the genus Micromonas -- which thrive in ecosystems ranging from tropical to polar -- look the same, but have evolved to contain different gene pools.
Bhattacharya said that understanding how these organisms change involves many issues.
The question, he said, is: "How do photosynthetic cells in the world's oceans recognize and adapt to their ever-changing environment and how will their latent abilities allow them to respond to climate change that will result in increased stratification and lower nutrient levels in the upper productive zone in oceans?
"To understand these complex issues, investigators need to generate gene catalogs from dominant plant organisms and understand how their genomes have evolved to thrive in vastly different oceanic regions ranging from near-shore to open ocean environments."
He said that the lead author of the Science article, Alexandra Z. Worden of the Monterey Bay Aquarium Research Institute and collaborators, addressed these key issues in oceanography by sequencing to completion the nuclear genome of two globally distributed, bacterial-sized green algae named Micromonas. One isolated sample (RCC299) came from tropical waters in the Pacific Ocean, whereas the other (CCMP1545) came from temperate Atlantic coastal waters off Plymouth, England.
"These picoeukaryotes are indistinguishable using cell morphology but turn out to be enormously different at the genome level," Bhattacharya said. "On average, these isolates share only 90 percent of the roughly 10,000 genes each contains, indicating they comprise distinct species. More remarkable is the finding of novel repeated sequences that have spread into genes of Atlantic sample that are completely missing in the Pacific sample."
He said that it is unclear how these ubiquitous elements originated or what their function might be in the Atlantic sample, but their presence demonstrates the distinct genomic trajectory that the two species have taken.
"Overall the genomes of these Micromonas species show clear indications of selection acting on the gene pool with each containing a set of unique genes acquired by horizontal gene transfer that are not shared with the other," he said. "These genes likely hold clues to how each species has adapted to its own specific marine environment."
"The work highlights the extent to which genomic diversity is hidden by a simple, shared morphology and points to the need to decipher gene functions in Micromonas to understand their role in adapting to regimes that define myriad marine environments," he said.
Source: University of Iowa (news : web)