Super-resolution microscopy takes on a third dimension
The shapes of some of the tiniest cellular structures are coming into sharper focus at the Howard Hughes Medical Institute's Janelia Farm Research Campus, where scientists have developed a new imaging technology that produces the best three-dimensional resolution ever seen with an optical microscope.
With this new tool, scientists can pinpoint fluorescent labels in their images to within 10-20 nanometers - about ten times the size of an average protein - in all three dimensions. The researchers say they now have an extremely powerful technology that will help reveal how biomolecules organize themselves into the structures and signaling complexes that drive cellular functions.
Their new method adds a third dimension to a cutting-edge form of light microscopy that scientists at Janelia Farm have used for the last two years to create two-dimensional images that pinpoint the location of fluorescently labeled proteins with extremely high resolution. To push this form of microscopy to the next level -- three-dimensional imaging -- the researchers borrowed a strategy widely used in industry to measure vanishingly small distances, such as the subtle variations in height on the surface of a computer chip.
Janelia Farm scientist Harald Hess and his colleagues adapted that technique, known as interferometry, to make it compatible with the fluorescent molecules often used by biologists to visualize proteins. When interferometry is combined with the super-high resolution photoactivated localization microscopy (PALM), researchers can see the three-dimensional architecture of cellular structures in extraordinary detail.
"This will be a good tool to really untangle things right down to the molecular structure level," said Hess, who led the development of the new technology in the applied physics and instrumentation group at Janelia Farm.
Hess and collaborators at the National Institutes of Health, Florida State University, and Janelia Farm, who call their new tool interferometric photoactivated localization microscopy (iPALM), have already created detailed images of three-dimensional structures previously not resolvable with light microscopy. Their "photo gallery" includes images of the microtubules that give cells structure; the two layers of a cell's outer membrane; and the focal adhesions that attach cells to their environment. Some of these images are included in a research article published in the February 2, 2009, issue of the Proceedings of the National Academy of Sciences describing the new technique.
Hess and Janelia Farm colleague Eric Betzig invented the PALM microscope in 2005. Scarcely three years later, it was one of a handful of new methods of "super-resolution" microscopy that were honored by Nature Methods in January 2009 as the "Method of the Year" for the previous year.
PALM permits biologists to visualize cells with far more detail than conventional optical microscopes, which are inherently limited by the wavelength of light. To achieve this resolution, PALM uses fluorescent labels that can be turned on and off with a pulse of light. Cells whose proteins are tagged with these labels are imaged repeatedly with PALM, with only a tiny subset of the fluorescent molecules turned on in each image. By compiling many thousands of these images, PALM creates a complete picture of the structure under study, pinpointing each fluorescently tagged protein. As a result, researchers get a much clearer picture than the overlapping haze that results when all of the tagged proteins are lit up at the same time, as in traditional fluorescence microscopy.
Hess and Betzig designed the concept for the first PALM microscope prior to their arrival at Janelia Farm in 2006. By the time they settled into their new laboratories, Hess says, both were already thinking about ways to improve the technique. Betzig focused on adapting the technique so that it could be used with living cells and with several differently colored fluorescent labels. For Hess, a major next goal was to expand the extraordinary spatial resolution they had already achieved to all three dimensions.
Hess, who spent 10 years working in the data storage and semiconductor industries, quickly focused his thinking on interferometry as a way to identify a protein's precise depth within a biological sample, and September of 2006 proposed the idea of iPALM. "Interferometry is one of the more sensitive measurement techniques out there," Hess said. "If you have bright enough light sources, you can measure ridiculously tiny displacements - way below the size of an atom."
When he worked in the hard disk industry, Hess used interferometry to detect subtle convolutions on the surface of a hard drive disk. The approach, he said, involved bouncing light off the surface of the disk and comparing the returned light wave to a "reference wave," which had been bounced off a mirror a known distance from the light source. "If light goes down and bounces off a surface, if that surface is a little bit higher or a little bit lower, that wave's going to be coming at you a little bit later or a little bit sooner," he explained. If the mirror and the experimental surface are the same distance from the light source, the waves, when added together, will cancel one another out. But tiny discrepancies in the two distances will shift the waves a measurable amount. "Depending on the amplitude of the summed waves," he said, "you can determine the vertical position to within nanometers."
No one had figured out how to apply the technique to biological samples, however. The primary challenge, Hess explained, was that in fluorescence microscopy, the key light waves travel from fluorescent tags within the sample itself, not from a readily manipulated laser. "It's a whole new paradigm," he said. "It isn't like you can go in there and take a piece of the laser to make a reference beam."
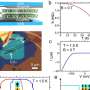
Hess and Janelia Farm colleague Gleb Shtengel saw a way around the problem: They decided to split each particle of light emitted from the fluorescent molecule in two. By splitting the photons, the researchers knew that each fluorescent photon would act as its own reference beam. They adapted the standard PALM microscope to collect this light both above and below the sample. Both of those beams of light travel to a custom-made beam-splitter, which divides the beam and sends it to three different cameras. A molecule's depth within the sample determines how much light reaches each of the cameras. "We record an image triplet, and depending on how much appears in camera one, two and three, we can say 'this was the height.' This is by far the most sensitive way of measuring vertical height," Hess said.
"iPALM needs only a modest amount of light to generate its sensitive measurements, and that's important for biological imaging," Hess says. Imaging techniques that demand more photons can force researchers to label the proteins they want to see with brighter dyes - which are often bulky and require harsh sample preparations that damage cells. Fluorescent probes such as those compatible with iPALM, on the other hand, can be genetically encoded so that they are manufactured by cells themselves. The power of these glowing markers was recognized with the 2008 Nobel Prize in Chemistry, which was awarded to HHMI investigator Roger Y. Tsien, Osamu Shimomura, and Martin Chalfie for the discovery and development of the first such tool, green fluorescent protein.
"It's beautiful if you can just have the cell attach the label for you," Hess said. "But the photoactivatable molecules that are used for fluorescence can only spill out so many photons before they finally fizzle. You have to make the most of what you've got. This technique is almost 100 times more efficient than other approaches [for three-dimensional imaging], in terms of harvesting the most information for the brightness of that fluorescent molecule."
Source: Howard Hughes Medical Institute