Self-digestion as a means of survival
In times of starvation, cells tighten their belts: they start to digest their own proteins and cellular organs. The process - known as autophagy - takes place in special organelles called autophagosomes. It is a strategy that simple yeast cells have developed as a means of survival when times get tough, and in the course of evolution, it has become a kind of self-cleaning process. In mammalian cells, autophagosomes are also responsible for getting rid of misfolded proteins, damaged organelles or disease-causing bacteria.
If this process malfunctions, it can result in infectious diseases, as well as cancer, Parkinson's or Alzheimer's disease. Biochemists at Frankfurt's Goethe University, working together with scientists from the University of Tromsø in Norway, the Weizmann Institute in Israel and the Tokyo Metropolitan Institute in Japan have just come up with an explanation as to how autophagosomes know exactly which proteins and organelles they should degrade.
"Although autophagy has been known for more than 30 years, it is astonishing that no-one thought of looking for the receptors that make this process so selective" explains Prof. Ivan Dikic from the Institute of Biochemistry II and the Cluster of Excellence 'Macromolecular Complexes' in Frankfurt. He had a head start in this field, since over several years, he and his group have researched and now published their work on another self-cleaning process in the cell: the degradation of small proteins in the proteasome, which acts as a kind of molecular shredder.
"We know that the molecules which are destined to be discarded are marked with the small protein ubiquitin and this is recognised by a receptor located at the gateway to the proteasome. It was natural to suggest a similar recognition mechanism for protein degradation by autophagosomes", says Dikic.
Unlike the proteasome, which is a complex molecular machine, autophagosomes simply consist of a double membrane that floats around in the cytoplasm. Not unlike white blood cells, they can engulf larger proteins or even whole cell organelles. But since they have no enzymes with which they can digest their own cargo, they fuse with lysosomes. When a Yoshinori Ohsumi's group in Japan reported that they had discovered ubiquitin-like proteins (ATG8) on the outer surface of the autophagosome and gone on to prove that they were specific for autophagy, Dikic and his colleague Dr. Vladimir Kirkin immediately began their search for potential autophagy receptors that might bind to the family of ATG8 proteins.
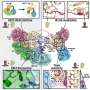
The team of international scientists report in the current issue of the renowned journal "Molecular Cell", that by employing methods from cell biology, biochemistry and mouse genetics, they have been able to identify a further protein, in addition to the known p62/SQSTM1 protein, that may act as a receptor. This is the protein NBR1, which has long been associated with cancer. Both proteins have a similar chain-like structure. At one end they bind to the ubiquitin that marks the protein aggregates and organelles that are to be degraded. Next to the ubiquitin-binding site is a domain that binds to the ATG8 proteins found at the autophagosomal membrane. Here, the protein waste can dock onto the autophagosome and can then be wrapped up in the membrane.
Vladimir Kirkin, who is now at Merck Serono in Darmstadt, is continuing these investigations with the long-term aim of developing new drugs. Dikic and his group are now concentrating on mitochondria - which are implicated in oxidative stress in cells - hoping to locate the receptors for autophagy on these important organelles.
More information: www.cell.com/molecular-cell/ab … 1097-2765(09)00064-1
Source: Goethe University Frankfurt