This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
proofread
Chiral detection of biomolecules based on reinforcement learning
As one of the basic physical properties, chirality plays an important role in many fields. Especially in biomedical chemistry, the discrimination of enantiomers is a very important research subject. Most biomolecules exhibit weak chirality in the ultraviolet band, so direct optical chiral detection is very difficult.
It is reported that surface plasmons enhance biomolecular circular dichroism. The coupling effect of metallic nanostructure and biomolecules is considered the basis of chiral detection. Finding appropriate nanostructures is crucial to achieving sensitive detection. Due to the complexity of the coupling model, it is difficult to do quantitative analysis and design nanostructures.
As one of the main artificial intelligence algorithms, reinforcement learning is inspired by behaviorist psychology, concerned with how software agents ought to take actions in an environment to maximize some notion of cumulative reward. The involvement of machine learning contributes significantly to nanophotonic development. Intelligent algorithms provide diverse optimization toolboxes to speed up prototyping of photonic device with enhanced performance.
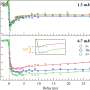
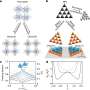
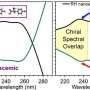
The research group of Prof. Zheyu Fang from Peking University introduced a nanostructure design platform based on reinforcement learning and the manipulation of optical properties. When biomolecules approach metallic nanostructures, their coupling effect modulates the circular dichroism spectrum and causes a shift of the resonance frequency. The algorithm successfully proposed nanostructures with high quality factor circular dichroism spectrum that emphasized the frequency shift. Microfluidic technology is involved to achieve highly sensitive chiral dynamic discrimination of glucose enantiomers. This method provides an example for the combination of artificial intelligence and optical sensing.
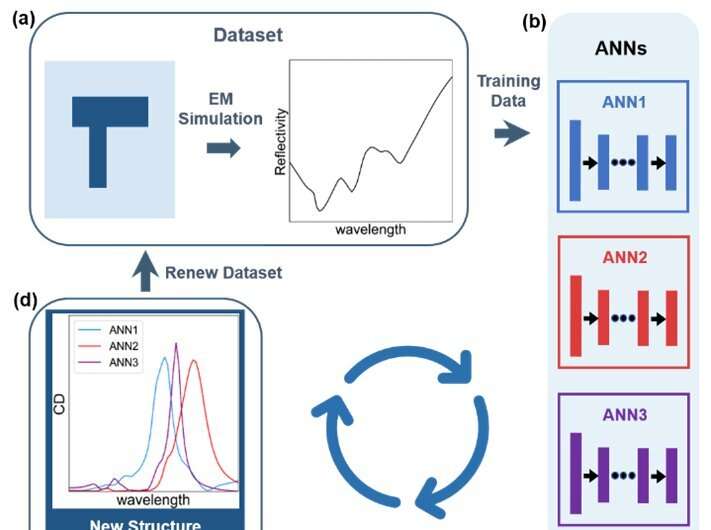
Compared with supervised learning, reinforcement learning significantly reduces the computational resources consumed by simulation. Sufficient electromagnetic simulations are the basis for supervised learning to predict accurately the optical response of various metal nanostructures. The training set of neural networks must contain all configurations, most of which possess little chirality. The simulation of achiral nanostructures is meaningless for ultimate optimization. On the contrary, the parameter exploration of reinforcement learning is simultaneous with model training. After several rounds of searching, the exploration range is limited in chiral nanostructures, so the algorithm does not waste computational resources on achiral nanostructures.
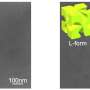
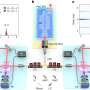
Metallic nanostructures are fabricated at the bottom of the microfluidic chip, and the chirality of the sample solution is monitored continuously through the real-time observation of spectral shifts. Unlike traditional biochemical methods, this detection realizes chiral discrimination without chemical reaction. Due to its low sample demand and low destructiveness, the method has great application value and the potential to achieve sensitive chiral detection for various biological macromolecules.
The paper is published in the journal Opto-Electronic Science.
More information: Yuxiang Chen et al, Chiral detection of biomolecules based on reinforcement learning, Opto-Electronic Science (2023). DOI: 10.29026/oes.2023.220019
Provided by Compuscript Ltd