June 22, 2017 report
Researchers identify mammals that are most likely to harbor viruses risky to humans
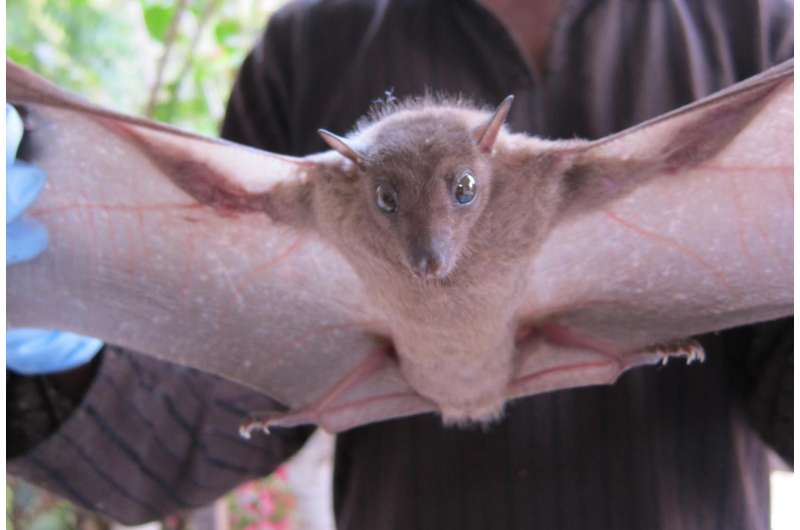

(Phys.org)—A team of researchers with the EcoHealth Alliance has narrowed down the list of animal species that may harbor viruses likely to jump to humans. In their paper published in the journal Nature, the group outlines the process they used to collect viral data on mammals around the globe, sorted them into groups and listed where they live. James Lloyd-Smith with the University of California offers a News & Views piece on the work done by the team in the same journal issue.
Scientists know that many of the viral threats we humans will face in the future are likely to come from viruses that already exist but reside in other species, particularly other mammals. The animal hosts have built up some degree of immunity to them, but we have not. Thus, if they jump to us, the result can be devastating. In this new effort, the researchers sought to catalogue all of the known viruses that infect mammals around the globe and identify which are most likely to jump to humans.
To create such a catalogue, the researchers created a database that held information on 754 mammal species, which represented 14 percent of all known mammals. They also added approximately 600 known viruses that infect mammals (of which a third were known to jump to humans) and which animals they infect. Next, they created mathematical models to use information in the database to provide useful information regarding the likelihood of a virus jumping to humans.
The researchers report that their models suggest that the likelihood of a virus jumping from a mammal species to humans depends heavily on species and geography. Bats were found to carry the largest number of viruses likely to jump to humans and the areas where it was most likely to occur were South and Central America. Primates posed the second largest risk factor, particularly in Central America, Africa and Southwest Asia. Rodents came in third with the risk most pronounced in North and South America and Central Africa.

Information gleaned from the system created by the researchers could prove more important over time as more data is added and the risk of a virus jumping rises. The hope is that it can be used to predict the next jump, allowing health officials time to prepare, or perhaps even to prevent it from happening.
More information: Kevin J. Olival et al. Host and viral traits predict zoonotic spillover from mammals, Nature (2017). DOI: 10.1038/nature22975
Abstract
The majority of human emerging infectious diseases are zoonotic, with viruses that originate in wild mammals of particular concern (for example, HIV, Ebola and SARS). Understanding patterns of viral diversity in wildlife and determinants of successful cross-species transmission, or spillover, are therefore key goals for pandemic surveillance programs. However, few analytical tools exist to identify which host species are likely to harbour the next human virus, or which viruses can cross species boundaries. Here we conduct a comprehensive analysis of mammalian host–virus relationships and show that both the total number of viruses that infect a given species and the proportion likely to be zoonotic are predictable. After controlling for research effort, the proportion of zoonotic viruses per species is predicted by phylogenetic relatedness to humans, host taxonomy and human population within a species range—which may reflect human–wildlife contact. We demonstrate that bats harbour a significantly higher proportion of zoonotic viruses than all other mammalian orders. We also identify the taxa and geographic regions with the largest estimated number of 'missing viruses' and 'missing zoonoses' and therefore of highest value for future surveillance. We then show that phylogenetic host breadth and other viral traits are significant predictors of zoonotic potential, providing a novel framework to assess if a newly discovered mammalian virus could infect people.
Journal information: Nature
© 2017 Phys.org