Using computer analysis, researcher seeks to heal vegetables affected by harmful fungi
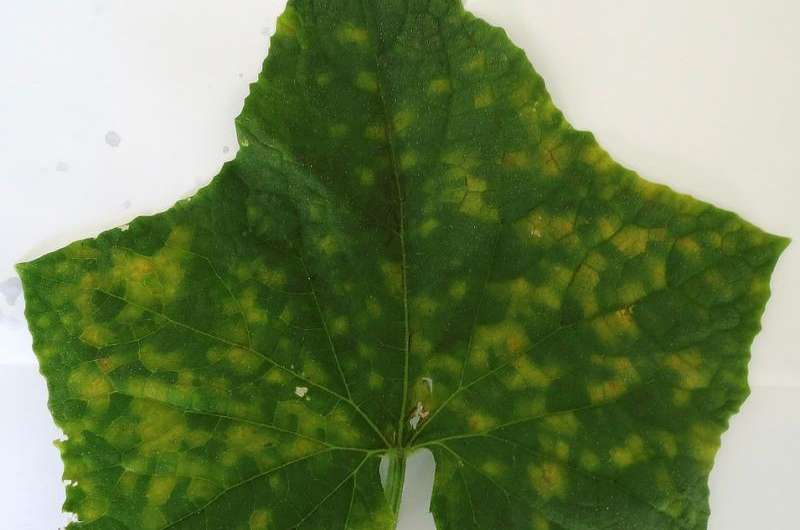

Crops such as cucumber, watermelon, squash and cantaloupe are affected by the fungi Pseudoperonospora humuli and Pseudoperonospora cubensis, both of which produce the downy mildew disease. Now, bioinformatics technology can identify genes of those pathogens in order to determine the ideal treatment.
Researchers want to understand at a genomic level the differences between these fungi. If genes are expressed in one fungus but not in another, they serve as markers that will indicate the appropriate treatment in these crops, says Dr. Elsa Gongora Castillo, plant biotechnologist who works at North Carolina State University.
She conducts computer analysis to identify fungi that affect the plant family Cucurbitaceae (cucumber, pumpkin, cantaloupe, watermelon).
"It's a bit like human disease, but in plants To understand the pathogen and its interaction with the plant allows development of a functional cure to treat the affected plants," says the specialist in plant genomics.
The researcher explains that she seeks to define alternative systems beyond conventional fungicides for cucumber, pumpkin, cantaloupe and watermelon.
The downy mildew disease affecting these plants is caused by a fungal infection; therefore, research aims to analyze their differences. First, samples of leaves from these plants are collected for in vitro cultures to isolate the fungi; then the DNA and RNA of fungi are extracted for sequencing, and through bioinformatic analysis, the researcher can determine the presence or absence of genes in the genomes of one species against the other.
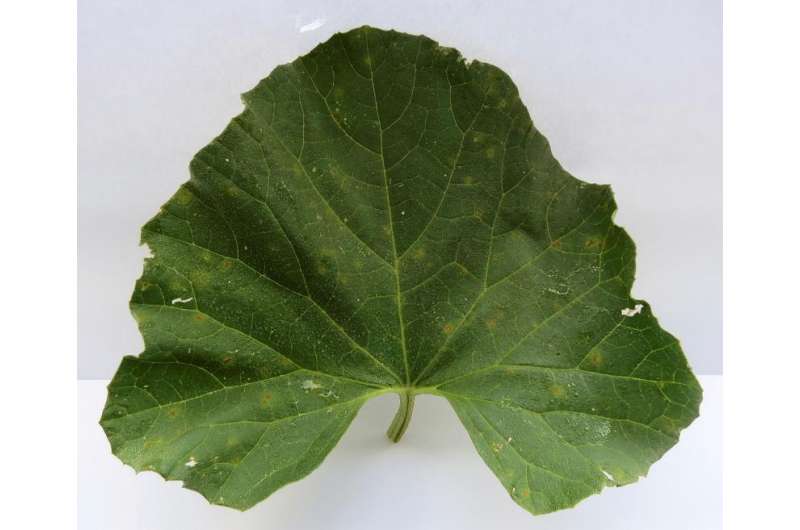

"With the sequences of pathogens, I identify the genes and their expression level; in other words, to study genes, I compare two of them—in this case, genes from Pseudoperonospora cubensis against Pseudoperonospora humuli. I see how many sequences there are for a given gene of P. cubensis and how many are there for the same gene P . humuli; if there is a difference in the number of sequences, we say that this gene is expressed or repressed; or if the gene is present or not in the genome," says Gongora.
Currently, laboratory results obtained from computer analysis are being corroborated and the research team seeks to benefit American farmers and their crops. This analysis will contribute to study models with which to understand other large-scale phenomena.
"Our model can be replicated elsewhere, in Mexico for example. But I think it takes a lot of investment in science to turn heads, and above all, a strong link between industry and science," says Gongora.
Provided by Investigación y Desarrollo