Blood reveals Great Barrier Reef sharks as homebodies

Small Australian sharks have been exposed as bigger homebodies than previously thought, in a study that took an existing chemical tracking technique and made it work for Great Barrier Reef sharks.
The study found that the travel history of the Australian sharpnose shark was written in their blood—with chemical 'fin-prints' showing they tended to stay within smaller areas than previously believed.
"Small-bodied sharks that are both predator and prey, such as the Australian sharpnose, may be particularly important links between food webs," says lead researcher Dr Sam Munroe, who studied the sharks while at James Cook University in Townsville.
"Information on their movements can improve our understanding of how the ecosystems function, while also helping us predict species most at risk from the impacts of a changing environment."
It was the first time the chemical tracking technique—known as stable isotope analysis—had been used to estimate shark movement over regional, coastal scales, and the results showed that they remained within the same 100km area for up to one year. The technique is often used to track land mammals, birds, and insects, and in marine animals (including sharks) over very large (e.g. whole oceans) or very small scales.
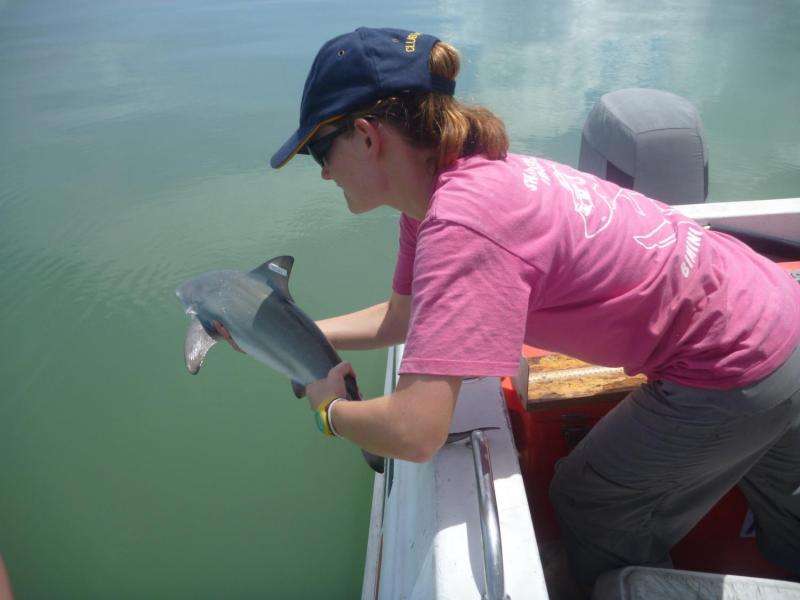

Different levels of elements such as carbon and nitrogen are naturally found in different environments. Sam compared these levels from seagrass and plankton in different bays along the coast with the levels in blood and muscle samples from the sharks, to map where the sharks had been.
"We didn't know if it would work for sharks on this scale," Sam says.
"But now we have shown it can be done for at least some shark species and we can hopefully apply this cost-effective approach in other environments."
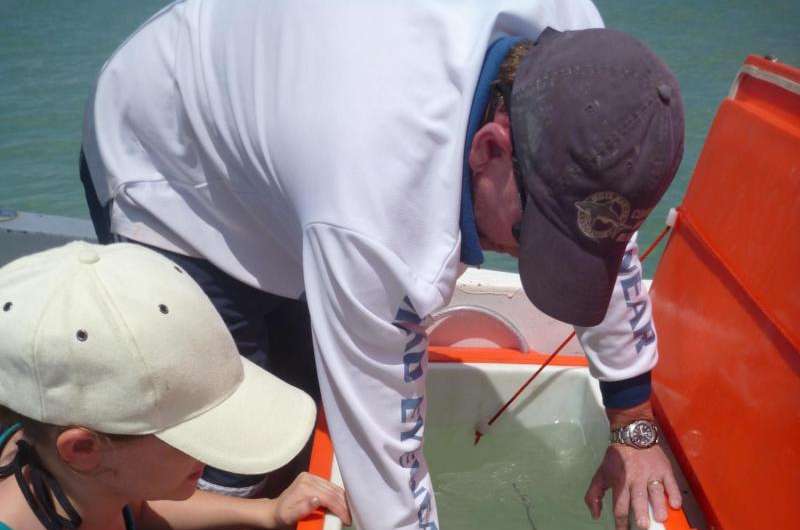

Satellite tagging is a popular tracking method and provides detailed movement information over huge scales, but can cost thousands of dollars per animal tagged. There is also the possibility of technical difficulties or of tags falling off, Sam says.
"Using isotopes means we don't get the same level of detail, but they only cost around $10–15 per tissue sample, and we get a broad understanding of where the animal was in the last year."
Sam says the small travel distances of the sharks may be due to a trade-off between moving widely to maximise their food and mating opportunities, or staying put to conserve energy.
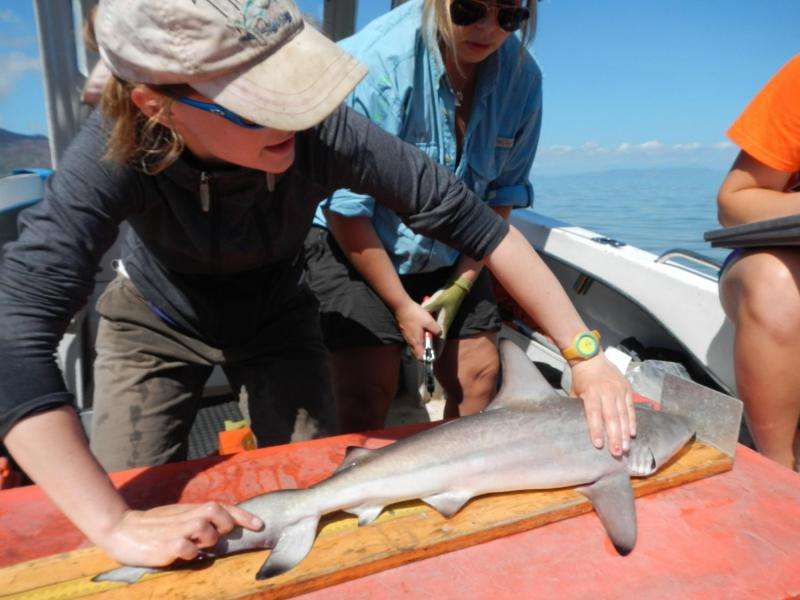

While the Australian sharpnose is abundant and not targeted by fishers, Sam hopes to see these tracking methods utilised for endangered species in the future.
Now pursuing post-doctoral studies at Griffith University in Brisbane, Sam's next step will be to apply these techniques to study the diet of deep-water sharks.
Provided by Science in Public