March 29, 2013 report
Researchers devise technique to allow X-ray crystallography of un-crystallized molecule groups
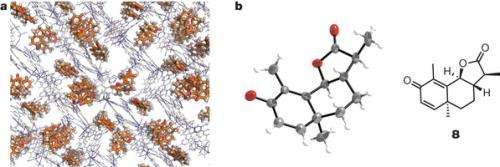
(Phys.org) —A team of researchers working in Japan has developed a method for allowing X-ray crystallography to work on molecular groups that have not first been crystallized. In their paper published in the journal Nature, the group describes how they built small scaffolds that resemble pockets for the molecules to rest in, securing them in place and allowing for X-ray crystallography analysis.
Over the years, X-ray crystallography has become one of the more important tools of modern research—it allows for the shape of molecules to be determined, which is important because molecular shape determines how different molecules can bind together. One roadblock to its use in some areas has been the difficulty in getting some molecule groups to crystallize—a necessary prelude to using X-ray crystallography because it causes the molecules to sit still in a way that allows them to be analyzed. In this new research effort, the team in Japan has found a way to allow researchers to use X-ray crystallography without having to get molecule groups to first crystallize.
They did it by constructing a scaffold of metal-organic frameworks to form octahedral cages, or pockets—they pull in molecules and hold them in a stable orientation which in turn allows them to be studied using X-ray crystallography. The team describes their structures as "crystalline sponges" that pull in small quantities of molecules and after a short stabilization period, hold them in place.
The researchers tested their new method by correctly determining the shape of several molecules whose shape was already known. One molecule shape in particular—miyakosyne A—was discerned by their method, which proved its feasibility as researchers have never been able to get it to crystallize and thus be studied using X-ray crystallography.
The only drawback seen thus far with the new technique is that it's limited to molecules of a certain size range—proteins for example, are made of molecules too large to fit in the cages the team developed. Because of that, the researchers are currently looking at ways to create larger cages and hope to one day develop a means of allowing for capturing and holding molecules of any size.
More information: X-ray analysis on the nanogram to microgram scale using porous complexes, Nature 495, 461–466 (28 March 2013) doi:10.1038/nature11990
Abstract
X-ray single-crystal diffraction (SCD) analysis has the intrinsic limitation that the target molecules must be obtained as single crystals. Here we report a protocol for SCD analysis that does not require the crystallization of the sample. In our method, tiny crystals of porous complexes are soaked in a solution of the target, such that the complexes can absorb the target molecules. Crystallographic analysis clearly determines the absorbed guest structures along with the host frameworks. Because the SCD analysis is carried out on only one tiny crystal of the complex, the required sample mass is of the nanogram–microgram order. We demonstrate that as little as about 80 nanograms of a sample is enough for the SCD analysis. In combination with high-performance liquid chromatography, our protocol allows the direct characterization of multiple fractions, establishing a prototypical means of liquid chromatography SCD analysis. Furthermore, we unambiguously determined the structure of a scarce marine natural product using only 5 micrograms of the compound.
Journal information: Nature
© 2013 Phys.org