Shared genes with Neanderthal relatives not unusual
During human evolution our ancestors mated with Neanderthals, but also with other related hominids. In this week's online edition of PNAS (Proceedings of the National Academy of Sciences), researchers from Uppsala University are publishing findings showing that people in East Asia share genetic material with Denisovans, who got the name from the cave in Siberia where they were first found.
Our study covers a larger part of the world than earlier studies, and it is clear that it is not as simple as we previously thought. Hybridization took place at several points in evolution, and the genetic traces of this can be found in several places in the world. We'll probably be uncovering more events like these, says Mattias Jakobsson, who conducted the study together with Pontus Skoglund.
Previous studies have found two separate hybridization events between so-called archaic humans (different from modern humans in both genetics and morphology) and the ancestors of modern humans after their emergence from Africa: hybridization between Neanderthals and the ancestors of modern humans outside of Africa and hybridization between Denisovans and the ancestors of indigenous Oceanians. The genetic difference between Neandertals and Denisovans is roughly as great as the maximal level of variation among us modern humans.
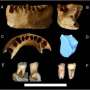
The Uppsala scientists' study demonstrates that hybridization also occurred on the East Asian mainland. The connection was discovered by using genotype data in order to obtain a larger data set. Complete genomes of modern humans are only available from some dozen individuals today, whereas genotype data is available from thousands of individuals. These genetic data can be compared with genome sequences from Neandertals and a Denisovan which have been determined from archeological material. Only a pinky finger and a tooth have been described from the latter.
Genotype data stems from genetic research where hundreds of thousands of genetic variants from test panels are gathered on a chip. However, this process leads to unusual variants not being included, which can lead to biases if the material is treated as if it consisted of complete genomes. Skoglund and Jakobsson used advanced computer simulations to determine what this source of error means for comparisons with archaic genes and have thereby been able to use genetic data from more than 1,500 modern humans from all over the world.
We found that individuals from mainly Southeast Asia have a higher proportion of Denisova-related genetic variants than people from other parts of the world, such as Europe, America, West and Central Asia, and Africa. The findings show that gene flow from archaic human groups also occurred on the Asian mainland, says Mattias Jakobsson.
While we can see that genetic material of archaic humans lives on to a greater extent than what was previously thought, we still know very little about the history of these groups and when their contacts with modern humans occurred, says Pontus Skoglund.
Because they find Denisova-related gene variants in Southeast Asia and Oceania, but not in Europe and America, the researchers suggest that hybridization with Denisova man took place about 20,000 years ago, but could also have occurred earlier. This is long after the branch that became modern humans split off from the branch that led to Neandertals and Denisovans some 300,000-500,000 years ago.
With more complete genomes from modern humans and more analyses of fossil material, it will be possible to describe our prehistory with considerably greater accuracy and richer detail, says Mattias Jakobsson.
Provided by Uppsala University