A challenge to improve Nuclear Magnetic Resonance for structural biology
(PhysOrg.com) -- In structural biology, the only technique available to predict the three dimensional structure of large complex molecules in solution, such as proteins and DNA, is NMR spectroscopy. To catalyze improvements in the techniques behind these predictions, the "eNMR" project has launched a new initiative. In September’s Nature Methods the project issued an invitation to the entire biomolecular Nuclear Magnetic Resonance community to participate in a large scale test of modern computing algorithms. This community-wide “contest” will potentially improve efficiency, reproducibility and reliability of NMR structure determination. eNMR will be using the Enabling Grids for E-sciencE infrastructure to power their analysis.
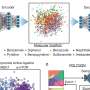
NMR spectroscopy is important in many different areas of science and is often used to determine the structure of complex molecules. The technique is particularly useful in biological sciences as it can predict the three dimensional structure of macromolecules in solution, including substances such as proteins and DNA that are key to understanding how the human body works. The analysis, however, is labour intensive and automation would accelerate the pace of research, helping scientists to identify molecules more quickly.
“Insight into the shape of biomolecules is the starting point for designing new drugs,” says Alexandre Bonvin, member of the eNMR project and one of the authors of the paper. “If we can improve this technology, it will help researchers in structural biology to be more productive. This could help shorten the whole process of designing new drugs.”
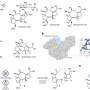
The small molecule ABT-737, for example, was found by screening a chemical library with NMR-based techniques. The discovery of ABT-737 was covered in the 2005 Nature paper “An inhibitor of Bcl-2 family proteins induces regression of solid tumours,” as a promising cancer fighting compound. (Though it has not, as of yet, been marketed.)
The eNMR project has worked to improve computational methods used for automation since late 2007, using EGEE’s computational resources to calculate molecular structures from NMR data. Their next step is to involve all interested stakeholders in their efforts. Through this challenge - called “Critical Assessment of automated Structure Determination of proteins by NMR” or CASD-NMR - the team invites laboratory researchers to submit molecules (technically the spatial coordinates of the atoms in the molecule with their associated NMR data) to help improve the algorithms used by the global eNMR team.
The CASD-NMR challenge will help computer scientists to automate NMR calculations and test them against blind datasets. The eNMR project and the National Institute of Health’s (NIH) Protein Structure Initiative are providing data for this challenge, and the CASD-NMR team hopes that other researchers will provide additional data sets.
In the future, automation in NMR will allow ‘unsupervised’ results to be accepted by the community as being correct and viable, ready for inclusion in the Protein Data Bank (PDB) straight away. The PDB is a database that stores macromolecular structural data that is freely and publicly available for further research (www.wwpdb.org).
“At this time fully automated methods are not reliable enough to be used blindly; this CASD-NMR experiment will be a valuable tool to see where we stand in automation and improve our methods,” says Bonvin.
CASD-NMR is set up to give the various teams eight weeks to apply automated methods to generate structures at a level of quality comparable to that of structures deposited into the PDB. National Grid Initiatives BigGrid in the Netherlands and IGI/INFN have contributed CPUs to the project so far. An assessment meeting is planned for mid-2010 to look at the results. Data are made available for CASD-NMR participants through the e-NMR project’s webpage (www.e-nmr.eu/CASD-NMR).
More information: CASD-NMR: critical assessment of automated structure determination by NMR, Nature Methods, Vol.6 No.9 September 2009 625, doi:10.1038/nmeth0909-625
Provided by CERN