New data suggest 'jumping genes' play a significant role in gene regulatory networks
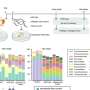
Research performed in the Center for Biomolecular Science & Engineering (CBSE) at the University of California, Santa Cruz, suggests that mobile repetitive elements--also known as transposons or "jumping genes"--do indeed affect the evolution of gene regulatory networks.
David Haussler, CBSE director and distinguished professor of biomolecular engineering at UCSC's Jack Baskin School of Engineering, said CBSE research teams are finding evidence that the early theories of Nobel Prize winner Barbara McClintock, later modeled by Roy Britten and Eric Davidson, are correct. Haussler will discuss these findings in a presentation on "Transposon-induced rewriting of vertebrate gene regulation" at the annual meeting of the American Association for the Advancement of Science (AAAS) in Chicago.
"Comparison of the human genome with the genomes of other species reveals that at least five percent of the human genome has been under negative selection during most of mammalian evolution," Haussler said. "We believe that this five percent is, therefore, likely to be functional."
Coding exons and structural RNA genes stand out because of their distinctive pattern of base substitutions and "indels"--the insertions and deletions of nucleic acid bases that can change the message in a genome. According to Haussler, however, most of the DNA under negative selection in vertebrate genomes does not appear to be transcribed and shares no sequence similarity with the genomes of invertebrates.
"Our research suggests that many of these elements serve as distal enhancers for developmental genes," Haussler said. "A significant amount of the gene regulatory material appears to have indeed been put into place by ancient transposons."
Source: University of California - Santa Cruz