Study specifies chemical pathway for ions through the cell membrane
(PhysOrg.com) -- Life ultimately depends on the traffic of tiny charged particles through porous proteins studding the membrane surrounding every cell. In research published in Nature, scientists at The Rockefeller University have for the first time mapped a stepping-stone pathway of amino acids that these charged particles, or ions, follow across cell membranes. Because the flow of ions underlies vital functions such as the electrical signaling of nerve, heart and muscle cells and the secretion of hormones and neurotransmitters, the work could have broad implications for understanding a range of diseases.
David C. Gadsby, the Patrick A. Gerschel Family Professor and head of the Laboratory of Cardiac and Membrane Physiology at Rockefeller, has for years researched the structure and function of a microscopic organic machine found in every animal cell called the sodium/potassium ion pump. This cellular gatekeeper pushes out of cells sodium ions that have leaked in and pulls back in potassium ions that have leaked out, maintaining a delicate balance that keeps cells alive and responsive to stimuli. The most recent advance in Gadsby’s research takes advantage of a technique that his lab pioneered, essentially turning this ion pump into a simpler type of membrane protein known as an ion channel.
While the mad rush of ions through an individual ion channel (millions per second) is large enough that the resulting current can be measured with an amplifying method called the patch-clamp technique, the current normally generated by a single ion pump (hundreds of ions per second) is far too weak to detect. Using a targeted lethal marine toxin, Gadsby, along with research associate Ayako Takeuchi and two former postdoctoral associates in the Gadsby lab, disabled the gates that control, and in doing so slow, the passage of ions through the pumps, unleashing a channel-strength current. “I like to borrow the techniques developed to study ion channels and use them to understand the pump,” Gadsby says. “The awesome power of patch-clamp recording is that you can follow the behavior of single molecules in their natural environment in real time.”
One by one, the researchers replaced the amino acids throughout the interior of the pumps with the amino acid cysteine. They exposed each variant to a highly reactive test ion a bit larger than the usual sodium and potassium ions that pass through the pump. Wherever the test ion could reach a cysteine, it reacted with it and remained attached, changing the flow of the current and allowing the researchers to find the exact path by identifying the amino acids that lined the space through which the ions passed.
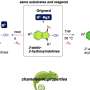
The researchers then mapped those amino acids onto a model of a related ion pump based on x-ray crystallography (see image, above) and found that the amino acids they had identified were contiguous and, moreover, surrounded by amino acids that did not react with the test ion at all. “The results agreed reasonably well,” Gadsby says, suggesting that the pathway traced through the sodium/potassium pump could be closely similar in related pumps that transport other ions, such as calcium.
“Individual ion binding sites have been proposed, but those are in a static structure,” says Takeuchi. “The patch-clamping method allows us to look at a real functioning pump that’s much closer to the dynamic physiological system. We’ve found the entire pathway and shown that it is consistent with the architecture of an ion pump from the same family.”
Article: Nature online: October 8, 2008 www.nature.com/nature/journal/ … abs/nature07350.html
Provided by Rockefeller University