Researchers use supercomputer to track pathways in myoglobin
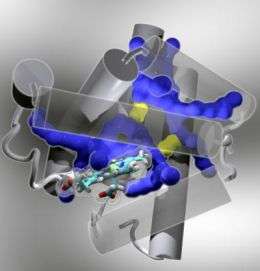
Some 50 years ago, after decades of effort, John Kendrew determined the structure of the small globular protein, myoglobin, which is responsible for oxygen storage in cells. For this discovery, he shared the Nobel Prize in chemistry with Max Perutz, who did similar work on hemoglobin. But a mystery remained: Exactly what paths does oxygen follow as it moves in and out of myoglobin?
An interdisciplinary team led by researchers at Virginia Tech has provided a computational solution to the decades-old puzzle.
Oxygen binds to an iron atom deep within myoglobin. How does the oxygen reach the iron through the solid protein wall? Thirty years of experimental and theoretical work have indicated that, as the protein fluctuates, transient "tunnels" may open up to form a network of pathways. But the exact nature and location of these dynamic pathways at atomic resolution have remained a mystery – until now, as reported in the online Early Edition of the Proceedings of the National Academy of Sciences (PNAS) the week of June 30, 2008.
"Every decade has seen breakthroughs in solving the puzzles this protein presented. Yet, exactly how small ligands such as oxygen or carbon monoxide (CO) migrate to the deeply buried binding site and out again was never completely resolved, in my opinion," said Alexey Onufriev of Blacksburg, Va., assistant professor of computer science and of physics at Virginia Tech.
He began his myoglobin research in 1997 as a postdoctoral researcher in chemistry faculty member Michael Prisant's lab at Duke University. According to Onufriev, Prisant suggested a computational geometry volume analysis. "That effort showed us what to look for," said Onufriev.
Ten years later, the Virginia Tech researchers proposed a multi-level computational solution to provide a detailed, complete, atom-level picture of ligand migration in myoglobin. "It ties together much previous experimental and theoretical understanding," said Onufriev.
The study is likely to be the first in which molecular dynamics succeeded to directly simulate the ligand migration all the way in and out of the protein at room temperature at atomic resolution.
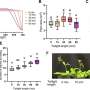
The simulations provided insights into several long-standing questions about myoglobin. They revealed that ligand migration occurs along two pathways with many branches and that ligands move into myoglobin using the same pathways that they use to move out of the protein. The simulations showed it is mainly local and short-lived fluctuations that are responsible for the opening and closing of these dynamic pathways. The simulations clearly show the pathways occur mostly between and around alpha helices – the protein scaffolds. Like people looking for an exit from an office complex, they cannot go through the walls or columns. Ligands pause at transient docking sites for 10s of nanoseconds. "Some of these sites were previously known but we found more," Onufriev said. "They are where pathways fork."
Another question was how the ligands moved. Do they force their way through the protein like a child in a play land ball pit or do they just go where the doors open? "Ligands follow the openings as they occur in the protein," Onufriev said.
Method
The work has also advanced methodology for the study of proteins. Myoglobin is the model protein – the guinea pig – of structural biology. "It has been thoroughly studied and has been key in elucidating many basic principles of the structure-to-function paradigm," said Onufriev.
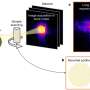
The group used tried-and-true molecular dynamic (MD) simulations complemented by a relatively simple, but robust computational geometry analysis of structural fluctuations available for ligand migration in myoglobin. "Our solution is on the edge of what is currently possible computation-wise, so Virginia Tech's SystemX supercomptuer was very helpful," Onufriev said.
The researchers used CO because it is easier to work with than oxygen and because most past experiments had also used CO. "The consensus is that small non-polar ligands such as oxygen, CO and nitric oxide follow basically the same pathways as oxygen ligands," Onufriev said.
One type of computational experiment initially puts ligands where they are bound, cuts the bond, and observes some of them wander out of the myoglobin protein. "Alternatively, we can place the ligand in water outside of the protein and occasionally observe it enter the protein and make its way all the way to the binding site," said Onufriev. "There is enough computational capacity in modern supercomputers to simulate enough of these events at room temperature to make statistically justified conclusions," he said.
"Another type of computation uses simple but highly-effective techniques from computational geometry and graph theory to look for space available for ligand migration based on free volume in the protein. For this one, we used snapshots from an MD simulation to find cavities in the myoglobin protein big enough to hold the ligand. Then we 'connected the dots' by computing the union of the cavities across all the snapshots and we clearly saw the pathways. These looked very similar to the ones we saw directly by following individual ligand trajectories," Onufriev said. "Over all, both methods complemented each other to provide a clear, simple picture of what was going on."
The methodology developed in this work can be used to study other systems. "There are many biomedically relevant systems that are much less understood, particularly as it concerns pathways, which are critical to protein function," said Onufriev.
Source: Virginia Tech