The key to unlocking the secret of highly specific DNAzyme catalysis
Using an extremely sensitive measurement technique, researchers at the University of Illinois have found clear evidence that a lead-specific DNAzyme uses the “lock and key” reaction mechanism. In the presence of zinc or magnesium, however, the same DNAzyme uses the “induced fit” reaction mechanism, similar to that used by ribozymes.
“The lock and key mechanism explains why this particular lead-specific DNAzyme makes such a sensitive and selective sensor,” said U. of I. chemistry professor Yi Lu, a corresponding author of a paper accepted for publication in Nature Chemical Biology, and posted on the journal’s Web site.
“Understanding the relationship between conformational change and reaction is important in obtaining deeper insight into how DNAzymes work and for designing more efficient sensors,” said Lu, who also is a researcher in the university’s department of biochemistry, the Beckman Institute, and the Center of Advanced Materials for the Purification of Water with Systems.
In the early 1980s, RNA molecules that can catalyze enzymatic reactions were discovered and named ribozymes. This discovery was followed by demonstrations in the 1990s that DNA also can act as enzymes, termed deoxyribozymes or DNAzymes.
With only four nucleotides as building blocks, versus 20 in proteins, nucleic acid enzymes may need to recruit cofactors (helper molecules) to perform some functions. Metal ions are a natural choice, and indeed most nucleic acid enzymes require metal ions for function under physiological conditions (and therefore are called metalloenzymes).
Metalloenzymes use various modes for functions for which metal-dependent conformational change (induced fit) is required in some cases but not in others (lock and key). In contrast, most ribozymes require conformation change that almost always precedes the enzyme reactions.
Using an extremely sensitive measurement technique called single-molecule fluorescence resonance energy transfer, Lu, physics professor Taekjip Ha and their research team studied the metal-dependent conformational change and cleavage activity of a particular lead-sensitive DNAzyme.
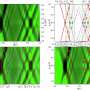
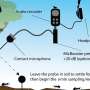
In single-molecule fluorescence resonance energy transfer, researchers add two dye molecules – one green and one red – to the molecule they want to study. Then they excite the green dye with a laser. Some of the energy moves from the green dye to the red dye, depending upon the distance between them.
“The changing ratio of the two intensities indicates the relative movement of the two dyes,” said Ha, who also is an affiliate of the university’s Institute for Genomic Biology and of the Howard Hughes Medical Institute. “By monitoring the brightness of the two dyes, we can measure conformational changes with nanometer precision.”
The researchers found that, in the presence of zinc or magnesium, a conformational change took place in the DNAzyme, followed by a cleavage reaction (behavior similar to many proteins and ribozymes). In the presence of lead, however, the cleavage reaction occurred without a preceding conformational change.
“This presents very strong evidence that the lead-specific enzyme uses the lock and key reaction mechanism,” Lu said. “This DNAzyme appears to be prearranged to accept lead for the activity.”
In previous work, Lu and his research group fashioned highly sensitive and selective fluorescent, colorimetric and magnetic resonance imaging sensors from the lead-specific DNAzyme used in this study. They have also constructed simple, disposable sensors using a different, uranium-specific DNAzyme.
“We think the answer to faster, more sensitive sensors lies with the lock and key mechanism,” Lu said. “Our next step is to look for other metals that use the lock and key mechanism with other specific DNAzymes. In addition, we want to investigate the structural details at the metal-binding sites and see how they change during catalysis.”
Source: University of Illinois at Urbana-Champaign