With no plan for DNA replication, cells depend on random selection
Each time a human cell divides it has to replicate three billion base pairs of DNA. All of the cell’s DNA must be copied once, but not more than once, within a very short period of time. But new research in yeast from Rockefeller University shows that instead of going about DNA replication in an organized way, the cell takes a very random approach, and the results leave scientists wondering how the cell ensures that the process is completed on time.
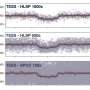
Bacteria, which have only 150,000 base pairs, have a single spot, called the origin, where replication begins. The origin is very easy to identify because the base pair sequence is always the same. But in higher organisms, from yeast to humans, origins are not so easy to identify, as there is no set sequence or position in the DNA. Researchers in Paul Nurse’s lab have completed the first genome-wide map of replication origins in the fission yeast Schizosaccharomyces pombe and measured their efficiency during both mitosis and meiosis.
“We wanted to know the origin positions, their usage and their distribution,” says Christian Heichinger, a postdoc in the Nurse lab and first author of the paper. “We found that origin distribution doesn’t match with their usage. We observed different large chromosomal domains enriched in either for efficient or inefficient origins.”
Even the most efficient origins aren’t used 100 percent of the time. In fission yeast, an efficient origin tends to be used only 30 to 50 percent of the time and weaker origins may only be used just 10 percent of the time. What’s more, there’s no way to predict which ones the cell will employ.
Scientists call this phenomenon the random completion problem: when origins appear to be selected randomly during replication, how does the cell ensure that all of its DNA gets replicated in time? “We don’t have the answer to this yet. It could be that S phase, the phase of the cell cycle in which DNA is replicated, is actually quite long, and that the cell extends this phase until replication is complete,” says Heichinger.
The researchers also found evidence that the position of the origin – in the context of the surrounding chromosome – may affect the origin’s efficiency. During meiosis, the process by which cell division of germ cells produces egg and sperm, the positions of the origins are similar to those used in mitosis, but their usage is different. “Origins in certain regions are induced, suggesting that the chromatin environment may have an effect on origin usage,” says Heichinger. “This hasn’t been shown before.”
By characterizing strong and weak origins, and noting their usage and distribution in yeast, scientists hope to glean clues to how replication is controlled in cells of higher organisms. “Without these exacting sequences, how is replication controlled? What goes wrong in diseases like cancer? This is our main interest, and this paper is just the beginning,” says Heichinger.
Ref: The EMBO Journal 25(21): 5171-5179 (November 2006)
Source: Rockefeller University