Viruses evolve to play by host rules
Biologists at the University of Pennsylvania and Harvard University have examined the complete genomes of viruses that infect the bacteria E. coli, P. aeruginosa and L. lactis and have found that many of these viral genomes exhibit codon bias, the tendency to preferentially encode a protein with a particular spelling.
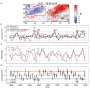
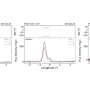
Researchers analyzed patterns of codon usage across 74 bacteriophages using the concept of a "genome landscape," a method of visualizing long-range patterns in a genome sequence.
Their findings extend the translational theory of codon bias to the viral kingdom, demonstrating that the viral genome is selected to obey the preferences of its host.
“The host bacterium is exerting a strong evolutionary pressure on the virus,” Joshua Plotkin, lead author and assistant professor in the Department of Biology at Penn, said. “This happens because a virus must hijack the machinery of its host in order to reproduce. We are seeing that viruses are forced to adopt the particular codon choices preferred by the bacterium they infect.”
The study found that each bacterium has a preferred way of spelling its genes. And it appears that viruses that infect a bacterium spell their own genes in the same way the bacterium does, obeying the rules of its host and demonstrating co-evolutionary behavior.
“Like a bee and a flower, an example of co-evolution between two large organisms, the same fundamental biological processes operate between two small organisms, as reflected in their genome sequences,” Plotkin said.
Moreover, the team found that the degree of codon bias varies across the viral genome. By comparing the observed genomes to randomly drawn genomes, the team demonstrated that the regions of high codon bias in these viral genomes often coincide with regions encoding structural proteins. Thus, the proteins that a virus needs to produce at high levels utilize the same encoding as its host organism does for highly expressed proteins.
Any protein can be encoded by multiple, synonymous spellings, but organisms typically prefer one spelling over others, a phenomenon known as codon bias. Codon bias is generally understood to result from selection for the synonymous spelling that maximizes the rate and accuracy of protein production.
The study, appearing in the current issue of the journal Public Library of Science Computational Biology, was performed by Plotkin and Grzegorz Kudla of the Department of Biology in the School of Afrts and Sciences at Penn and Julius Lucks and David Nelson of Harvard University.
Source: University of Pennsylvania