Breaking down Huntington's disease one protein at a time
Hoping to piece together the intricate series of interactions that lead to Huntington's disease, Indiana University Bloomington scientists have determined the shape and structure of a binding site that may prove useful in combating the neurodegenerative disease.
In the Feb. 1 issue of Journal of Molecular Biology, IU Bloomington biologists Joel Ybe and Qian Niu describe a region on the surface of HIP1 (Huntingtin-interacting protein 1) that could bind HIPPI (HIP1-protein interactor). The association of HIP1 and HIPPI is believed to lead to the degeneration of nerve cells.
"If we now think that this is the region where HIPPI binds, we could eventually design a drug that can come in and sit down between these two proteins, which would prevent the binding of HIPPI," said Ybe, who led the research.
Ybe and Niu's paper is the first to scrutinize a Huntington's disease-related protein's structure and function at the molecular level. Ybe and colleagues hope meticulous study of each Huntington's disease protein will suggest new avenues for wholesale prevention.
"The important thing for us is to come up with something that will potentially help people," said Ybe. "What is happening before the manifestation of the disease? Can we use this information to come up with drugs to diffuse that process?"
Huntington's disease is a hereditary disorder that causes large numbers of nerve cells to die. About 30,000 people in the U.S. are estimated to have the disease -- approximately one person in ten thousand. Symptoms include uncontrolled movements, dementia and depression, but these symptoms do not usually appear until the afflicted reach their 30s or 40s. Despite major strides forward in understanding the disease in recent years, there is currently no cure.
The disease begins when the huntingtin protein falls off HIP1. The vacancy allows another protein, HIPPI, to then bind to HIP1. The complex of HIP1 and HIPPI is responsible for activating other proteins that cause the death of cells. The loss of large amounts of nerve cells leads to a loss of motor function, and eventually brain function, too.
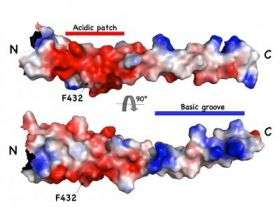
Ybe and Niu used X-ray crystallography to look at an area of interest on the surface of HIP1, which works in concert with clathrin to traffic nutrients into a cell, and has long been implicated as playing an important role in the development of Huntington's disease. They learned that the potential binding surface of HIPPI in HIP1 has an unexpected shape for a binding site, a spiraling spiral called a "coiled coil." This finding was contrary to predicted results that the binding surface that receives HIPPI is folded into a so-called death effector domain.
Using the information from the published molecular structure of HIP1, IU biologists hope to be able to test which protein connections are ultimately responsible for triggering the chain of interactions leading to Huntington's disease and how to block them. For example, they observed that clathrin, protein involved in bringing nutrients to the cell, binds with HIP1 right next door to where HIPPI binds. While clathrin "packages" nutrients for a cell, HIP1 connects these "baskets" to the structure of the cell. If HIPPI binding with HIP1 prevents clathrin connection with HIP1, then the normal pathway of nutrients into a cell is interrupted, causing changes in the cells ability to function normally.
"Until we understand the relationship between huntingtin protein, HIP1, clathrin and HIPPI -- we are not going to understand what is happening in the person who has the disease," says Ybe. "You understand what's going on in healthy cells, before you understand what's going on in diseased cells."
Source: Indiana University















