Repeating genes
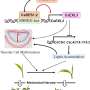
Huntington’s disease is a genetic time bomb: Programmed in the genes, it appears at a predictable age in adulthood, causing a progressive decline in mental and neurological function and finally death. There is, to date, no cure. Huntington’s, and a number of diseases like it, collectively known as trinucleotide repeat diseases, are caused by an unusual genetic mutation: A three-letter piece of gene code is repeated over and over in one gene.
Scientists at the Weizmann Institute have now proposed a mechanism that provides an explanation for the remarkable precision of the time bomb in these diseases. This explanation may, in the future, point researchers in the direction of a possible prevention or cure.
The number of repeats in Huntington’s patients ranges from 40 to over 70. Scientists have noted that, like clockwork, one can predict by how many times the sequence repeats in a patient’s gene both the age at which the disease will appear and how quickly the disease will progress. The basic assumption has been that the protein fragment containing the amino acid (glutamine) encoded in the repeating triplet slowly builds up in the cells until eventually reaching toxic levels.
This theory, unfortunately, fails to explain some of the clinical data. For instance, it doesn’t explain why patients with two copies of the Huntington’s gene don’t exhibit symptoms earlier than those with a single copy. Plus, glutamine is produced in only some trinucleotide diseases, whereas the correlation between sequence length and onset age follows the same general curve in all of them, implying a common mechanism not tied to glutamine.
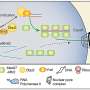
Research student Shai Kaplan in Prof. Ehud Shapiro’s lab in the Biological Chemistry, and Computer Sciences and Applied Mathematics Departments, realized the answer might lie in somatic mutations – changes in the number of DNA repeats that build up in our cells throughout our lives. The longer the sequence, the greater the chance of additional mutation, and the scientists realized that the genes carrying the disease code might be accumulating more and more DNA repeats over time, until some critical threshold is crossed.
Based on the literature on some 20 known trinucleotide repeat diseases and their knowledge of the mechanisms governing somatic mutation, Shapiro, Kaplan (who is also in the Molecular Cell Biology Department) and Dr. Shalev Itzkovitz created a computer simulation that could take a given number of genetic repeats and show both the age of onset and the way in which the disease progresses. Their findings appeared in PLoS Computational Biology.
The new disease model appears to fit all of the facts and to provide a good explanation for the onset and progression of all of the known trinucleotide repeat diseases. Experimentation in research labs could test this model, say the scientists and, as it predicts that all these diseases operate by somatic expansion of a trinucleotide repeat, it also suggests that a cure for all might be found in a drug or treatment that slows down the expansion process.
Source: Weizmann Institute of Science