Molecule stops DNA replication in its tracks
(PhysOrg.com) -- When a dividing cell duplicates its genetic material, a molecular machine called a sliding clamp travels along the DNA double helix, tethering the proteins that perform the replication. Researchers from the laboratory of Rockefeller University’s Michael O'Donnell, a Howard Hughes Medical Institute investigator, have discovered a small molecule that stops the sliding clamp in its tracks. The finding will enable scientists to better study the proteins that duplicate DNA, and may ultimately provide a platform for developing improved antibiotics.
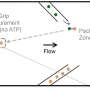
The process is akin to unzipping a zipper: The sliding clamp works its way along the DNA double helix while a network of proteins work together to unwind the two strands. Proteins known as polymerases then add, in assembly-line fashion, nucleotide bases — the building blocks that make up DNA — to convert the now-single-stranded templates into two new duplex DNA molecules. Bacteria have five known DNA polymerases (higher organisms such as humans have more), only one of which, polymerase III (pol III) is responsible for replicating the chromosome, while the others appear to be involved in DNA repair.
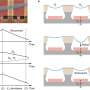
To better understand the functions of the other polymerases, O’Donnell and colleagues at Rockefeller used a combination of biochemical techniques to identify a small molecule that would inhibit the binding of the polymerases to the beta sliding clamp. With the help of researchers in Rockefeller’s High Throughput Screening Resource Center, coauthors Roxana E. Georgescu and Olga Yurieva, research associates in O’Donnell’s lab, screened some 30,000 compounds using a technique called fluorescence anisotropy. Georgescu and Yurieva looked for compounds that would disrupt the interaction of a fluorophore-labeled peptide with the peptide-binding pocket of the sliding clamp. Because the peptide is small and the clamp is big, the signal generated by the fluorophore is very different.
Georgescu and Yurieva identified one compound, called RU7, that differentially inhibited polymerases II, III and IV. RU7 did not inhibit pol IV at all, while pol III was inhibited the most.
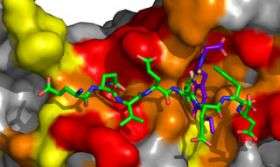
The researchers then used x-ray crystallography to compare how RU7 and polymerases II, III and IV bind to the clamp. They found that while all three polymerases and RU7 bind to the same peptide-binding site of the clamp, they do so in different ways. Polymerase IV, for example, forms additional contacts to the clamp outside of the peptide-binding site, which may account for its resistance to disruption by RU7.
“The role of polymerase III in replication has been very well studied, but the roles of the other polymerases are not well understood,” says O’Donnell. “RU7 may be an important tool as a chemical probe to better understand the functions of polymerases II and IV in normal cell growth and in response to DNA damage.”
Because RU7 halts the replication of bacterial DNA by disrupting polymerase III — but does not affect DNA replication in yeast, which uses the same molecular machinery as humans — O’Donnell’s research suggests that RU7 could provide a starting point for antibiotic drug design. Further tweaking, for example by adding atoms that enable the compound to fit into a second binding site, could even increase RU7’s potency.
Citation: Proceedings of the National Academy of Sciences 105(32): 11116–11121 (August 12, 2008)
hwmaint.pnas.org/cgi/content/full/105/32/11116
Provided by Rockefeller University