Multi-lab collaboration yields first detailed map of nuclear pore complex
A cell’s membrane-bound nucleus contains precious contents — its DNA — so it must be very careful about what enters and leaves this important space. To do this, it uses hundreds to thousands of nuclear pores as its gatekeepers, selective membrane channels that are responsible for regulating the material that goes to and from a cell’s DNA and the signals that tell a cell what to do and how to do it.
But the structure of each of these nuclear pores is so large, and so flexible, that it couldn’t be visualized in detail using existing methods. Now, in two landmark studies published in Nature, Rockefeller scientists have nailed down the first complete molecular picture of this huge, 450-protein pore and their findings provide a glimpse into how the nucleus itself first evolved.
The structure is the culmination of a three-laboratory, nine-year collaboration among Rockefeller professors Michael Rout and Brian Chait, and Andrej Sali, now at the University of California, San Francisco. Rout spearheaded the biochemical studies, Chait the mass spectrometry-based identification and characterization of the proteins and protein complexes and Sali was in charge of creating computational techniques that could take all this data and put it into countless configurations until it found the ones that fit. Together with a dedicated team of postdoctoral researchers, students and technicians, the group gathered and analyzed massive amounts of data to come up with a rough draft of the structure of the nuclear pore.
The computer sorted through about 200,000 different configurations of the pore’s component proteins, finding about 1,000 possible, very similar structures that fully satisfied all the thousands of restraints provided by experimental data: restraints such as which protein could be next to which, or where that large protein had to be in relation to this small one. Rout describes it as being a bit like a crossword puzzle that gradually comes together as each clue is solved. Their solution, he says, lets them see the position of every protein.
The scientists’ results have given them a peek into the early evolution of eukaryotic cells, the cells that make up all higher organisms, from people to plants to fungi. These cells split off from their progenitors when they developed a nucleus and other specialized organelles that allowed them to compartmentalize different aspects of cellular metabolism. And that compartmentalization was made possible by membranes and coating complexes, which act “like little hands,” Rout says, to grab, shape and stabilize membranes.
From what he and his colleagues can tell by looking at the nuclear pore, the sophisticated nuclear pores we have today arose directly from these early structures. “We think that once the cells gained this coating complex, they ran with it and started to duplicate it and specialize it,” he says. Different versions of this complex are apparent in other cell membranes, such as the endoplasmic reticulum and the Golgi, and likely plays analogous roles in all of them: grabbing the membrane, curving it and helping it move around.
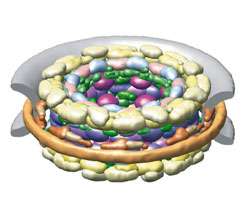
“Evolution is a process of duplication and divergence,” Rout says. He and his colleagues saw clear evidence of this when they color-coded the proteins in the pore. One method of color coding revealed alternating stripes, like spokes on a wheel: For every protein, there was another one that looked quite similar. Color coding a different way showed the same pattern in the pore’s outer and inner rings, one of which appears to be a slightly modified duplication of the other. These are evidence of duplication events, Rout says, showing that the evolution of the complicated nuclear pore was a more straightforward affair than previously thought. “It’s made of many different variations of a theme of just one unit.”
Every piece of the team’s complex structure matches evidence gleaned from electron microscope photographs and other, cruder models. And yet, despite its detail, it’s still just a draft. “It’s a map that’s given us a teeny bit of insight into how the nuclear pore functions, but we want to do much more work,” Rout says. “The nuclear pore is the communication device that the nucleus uses to communicate with the rest of the cell. And if you don’t understand how that works, you don’t understand a key part of how the cell works. And if you don’t understand that, you can’t completely understand how cancers work or how a single cell turns into a human being, or how a single grain of wheat turns into a whole crop. You have to see the cell as a machine and understand all of its parts.”
Nine years later, Rout, Chait and Sali are much closer to that understanding. And to assist others in getting there, they’ve helped build the NIH-funded National Center for Dynamic Interactome Research, headed by Rout, to guide other scientists through the process. With the center’s backing, the researchers hope that eventually new molecular puzzles should take only a year or so — rather than a decade — to solve.
Citations:
Nature 450(7170): 683–694 (November 29, 2007)
Nature 450(7170): 695–701 (November 29, 2007)
Source: Rockefeller University


















