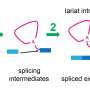
Adenine ‘Tails’ Make Tailored Anchors for DNA
Researchers from the National Institute of Standards and Technology, the Naval Research Laboratory and the University of Maryland have demonstrated a deceptively simple technique for chemically bonding single strands of DNA to gold.
Among other features, they report in a forthcoming issue of the Proceedings of the National Academy of Sciences, the technique offers a convenient way to control the density of the DNA strands on the substrate, which could be important for optimizing DNA sensor arrays.
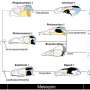
Short DNA sequences arrayed on substrates like glass, silicon or gold are used in biochemical sensors that can detect specific “target” sequences of DNA or analyze complex sequences. In such arrays, DNA strands are attached to the substrate by one end and stand up like bristles on a brush.
Specific “target” DNA sequences from a test sample can be identified because they will bond (hybridize) only to a complementary sequence on the array—microarray “gene chips” are the best-known example of the technology. The properties of gold are well-known, so it is a practicaland convenient substrate for some sensors. One popular technique for making DNA sensors (developed at NIST) is to use DNA with a sulfur atom attached to one end, which acts as “glue” because sulfur readily reacts with gold.
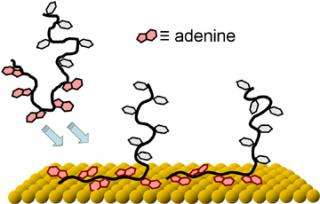
But a potentially less expensive and even simpler approach, according to the NIST, NRL and UMD team, might be to use a string of adenine nucleotides as an anchor. Of the four nucleotides that comprise DNA molecules, adenine, turns out to have a particularly high affinity for gold. Short strands composed entirely of adenine will adhere to a gold surface even if they have to muscle aside other strands in the process.
As a result, say the researchers, short blocks of adenine at the end of DNA strands can serve as bonding anchors—but even better, they say, these adenine blocks can be used to control the spacing of the DNA strands on the substrate. This is because each adenine tail lies flat on the substrate, taking up space. Within limits, the longer the adenine tail is, the larger is its footprint on the substrate, and the lower the total density of DNA strands.
Controlling the density of DNA “brushes” on a substrate is important for sensor design because an overly dense thicket does not leave enough room for “target” DNA strands from the test sample to bond, while too sparse an array doesn’t produce a strong enough signal.
The authors have applied for a patent on the technique.
Citation: A. Opdahl, D. Petrovykh, H. Kimura-Suda, M.J. Tarlov and L.J. Whitman. Independent control of grafting density and conformation of single-stranded DNA brushes. Proceedings of the National Academy of Sciences, 104:9-14 (2007) (Appears on-line in PNAS Early Edition.)
Source: NIST