Structural study backs new model for the nuclear pore complex

(PhysOrg.com) -- In higher organisms, the genetic material is confined and protected in the cell nucleus. In order for a healthy cell to function, the DNA must send manufacturing orders through the double membrane of the nucleus and into the cell’s cytoplasm, where the protein production factories are and where most cellular functions are carried out. The sole portals through which these instructions pass — nuclear pore complexes — have a say in what the orders are and how they are conveyed. But these conspicuously large structures have ironically proved all but inscrutable to researchers over the years.
“The nuclear pore complex is one of the most mysterious things in cell biology. It’s basically a black box,” says André Hoelz, a research associate at The Rockefeller University who studies the complex in John D. Rockefeller Jr. Professor Günter Blobel’s Laboratory of Cell Biology.
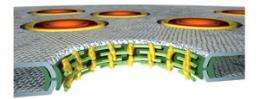
But more and more light is shining in. On the cover of the December 26 issue of Molecular Cell, available online today, Hoelz, Blobel and their colleagues illuminate the atomic structure of a key protein at the heart of the complex. The finding advances a new model that the lab proposed last year. The model proposes a cylindrical nuclear pore complex comprised of rings of alternating protein structures that zip together, providing the flexibility and space the pore needs to let pass a variety of materials in and out of the cell’s nucleus. “We were very excited to find a protein that is so similar in structure to what our model predicts,” Hoelz says.
For the past few years, Hoelz and his colleagues have been chipping away at the nuclear pore complex one protein at a time. Their piecemeal approach, using x-ray crystallography to identify the atoms in each protein, is necessary because the complex is too large and pliable to succumb to traditional imaging or genetic analysis, Hoelz says. The nuclear pore complex is built from about 30 different proteins and is 30 times the size of the ribosome, the site of protein production, and 1,000 times the size of the common ion channels found in all cell membranes. So far, the Blobel lab has solved the structure of four of seven proteins thought to provide a cylindrical coat for the pore membrane. They are about to publish the molecular makeup of a fifth, Hoelz says.
The structure to be published on Friday, a pair of nucleoporin proteins called Seh1-Nup85, is similar in shape and size to the last one the lab discovered, Sec13-Nup145C. Both pairs form bent rods and are about the same length as the depth of the nuclear pore complex found in yeast. This fits with the lab’s model, which suggests that these two protein pairs alternate around the interior lining of the complex, forming a picket fence of sorts. The model predicts that the remaining three proteins of this coat will horizontally join these protein pairs together. “Clearly, more work is required to know for sure, whether or not our model is correct,” says Hoelz.
Once researchers decipher the rest of the complex’s atomic structure, they will be able to precisely manipulate different parts and study the effects, hashing out the role this key portal plays in the process of turning DNA into proteins. “I think if we figure out the whole structure, we will open up a whole new field,” Hoelz says.
Given the central role of the nuclear pore complex in the most basic cell processes, defects in its assembly, structure and function can have lethal consequences. Its proteins have been associated with primary biliary cirrhosis, cancer, viral infections and triple A syndrome, says Erik Debler, postdoctoral fellow in the Blobel laboratory and first author of the paper. A better understanding of how the complex works could lead to treatments for these diseases. The nuclear pore complex is also of basic scientific interest as a key element in a very ancient evolutionary coup that is found in every cell more complicated than the simplest single-celled microorganisms: the nucleus.
“What distinguishes us from bacteria is that we have a cell nucleus,” Hoelz says. “It protects the DNA from environmental factors. It divides and segregates functions in different parts of the cell. It introduces all kinds of complexities that lower organisms do not have. The nuclear pore complex plays a big part in all of that.”
Paper: Molecular Cell (December 26, 2008) www.cell.com/molecular-cell/ab … 1097-2765(08)00840-X
Provided by Rockefeller University